Impact statement
The contribution of host genetic factors to COVID-19 has been investigated through genome-wide association studies (GWAS), as genetic targets generally double the success rate of drugs in clinical development. Researchers from around the world have teamed up and found 25 susceptibility loci, primarily related to immune response, but also to the ABO blood group system (with the most commonly replicated locus, the ABO gene), that influence either susceptibility to infection and/or progression of COVID-19. These genetic discoveries can now help suggest specific targets for drug reuse and new drug development. They also increase our knowledge of COVID-19 biology and our understanding of genetic risk factors for SARS-CoV-2 transmission. Some of the genes, for example, IFNAR2 and ACE2, encode proteins for which drug candidates are currently being tested in clinical trials. Strikingly, almost all independent (genetic and nongenetic) study data suggest that blood group A is associated with a higher probability of SARS-CoV-2 infection and blood group O with a lower probability, which is also reflected in the increased and decreased numbers of severely ill COVID-19 patients with blood group A and O, respectively. Model calculations show that the relative probability of SARS-CoV-2 transmission between an infected index person and an ABO-incompatible contact is reduced by an average of 40% in certain situations (ranging from 20 and 55% depending on ABO blood group frequency, estimated risk effect in different countries and the proportion of the population infected), which is confirmed by independent studies in couples. Current GWAS studies on refined COVID-19 symptoms such as loss of smell or taste and in the context of different demographic characteristics, but also in the context of recent data on reinfection with new viral variants and the human microbiome, are likely to help elucidate the underlying biology and provide precise and personalized treatments.
Introduction
The COVID-19 pandemic is a global crisis that has caused severe disruption in health systems and the global economy. Tremendous efforts have been made to contain the spread of SARS-CoV-2 and immunize the world’s population against the virus. At the same time, extensive research has been conducted to identify risk factors to protect vulnerable groups and to find new targets for drug development for severely ill individuals. To this end, a series of genome-wide association studies (GWAS) and genome-wide meta-analyses (GWMA) have been conducted in large patient populations to determine genetic determinants in the human genome of susceptibility to infection and disease severity in COVID-19. After two early blood group studies from hospitals in Wuhan and Shenzhen in China (which both first appeared on a pre-print server in March 2020 and were later published in peer-reviewed journals (Zhang et al., Reference Zhang, Huang, Xia, Fan, Zhu, Zhu, Zhang, Tao, Cheng and Chen2020; Zhao et al., Reference Zhao, Yang, Huang, Li, Gu, Lu, Zhang, Liu, Liu, Liu, He, Sun, Wei, Yang, Wang, Zhang, Zhou, Xing and Wang2021)) showed a statistical correlation of ABO blood group distributions with the incidence of COVID-19 compared to uninfected controls, nearly all (hypothesis-free) COVID-19 GWAS and GWMA reported associations between genetic variants at the ABO gene locus and ABO blood group distribution and susceptibility to SARS-CoV-2 infection and/or severity of COVID-19. The statistical observation that ABO blood groups and thus blood group antigens, in general, can increase or decrease host susceptibility to infection, including viral infections with norovirus, rotavirus, HIV, SARS-CoV-1 and influenza, is not a new finding, although the underlying mechanisms are not clearly known (Cooling, Reference Cooling2015). Anthropological studies indicated that the geographical distribution of human blood groups also reflects the susceptibility of populations with certain blood groups to plague, cholera, smallpox, malaria and other infectious diseases (Berger et al., Reference Berger, Young and Edberg1989). A well-known example is the increased host resistance to Plasmodium falciparum infection (malaria) in blood group 0 carriers in many African and Asian populations (Degarege et al., Reference Degarege, Gebrezgi, Ibanez, Wahlgren and Madhivanan2019), and it is estimated that 25% of the risk for malaria severity in Africa is determined by genetic factors of the human genome (Mackinnon et al., Reference Mackinnon, Mwangi, Snow, Marsh and Williams2005). In the following sections, I summarize which susceptibility loci for SARS-CoV-2 infection and/or COVID-19 disease severity have been identified to date through large-scale genome-wide analyses and what the genetic variants at the ABO gene locus and the ABO blood group associations might have to do with SARS-CoV-2 infection or severity in COVID-19. I will then review the current models of SARS-CoV-2 transmission in the context of ABO blood groups for validity based on study results and briefly look at the distribution of ABO blood groups in different populations worldwide to illustrate, what impact the statistical finding that blood group O is protective against infection and/or the severity of COVID-19 disease might have on the transmissibility of SARS-CoV-2 at the population level and at the individual level, and when blood group O might offer a real advantage over the other blood groups.
COVID-19 susceptibility loci from genome-wide studies
Of the 25 susceptibility loci with genome-wide significance identified to date in the human genome (Ellinghaus et al., Reference Ellinghaus, Degenhardt, Bujanda, Buti, Albillos, Invernizzi, Fernandez, Prati, Baselli, Asselta, Grimsrud, Milani, Aziz, Kassens, May, Wendorff, Wienbrandt, Uellendahl-Werth, Zheng, Yi, de Pablo, Chercoles, Palom, Garcia-Fernandez, Rodriguez-Frias, Zanella, Bandera, Protti, Aghemo, Lleo, Biondi, Caballero-Garralda, Gori, Tanck, Carreras Nolla, Latiano, Fracanzani, Peschuck, Julia, Pesenti, Voza, Jimenez, Mateos, Jimenez, Quereda, Paccapelo, Gassner, Angelini, Cea, Solier, Pestana, Muniz-Diaz, Sandoval, Paraboschi, Navas, Garcia Sanchez, Ceriotti, Martinelli-Boneschi, Peyvandi, Blasi, Tellez, Blanco-Grau, Hemmrich-Stanisak, Grasselli, Costantino, Cardamone, Foti, Aneli, Kurihara, ElAbd, My, Galvan-Femenia, Martin, Erdmann, Ferrusquia-Acosta, Garcia-Etxebarria, Izquierdo-Sanchez, Bettini, Sumoy, Terranova, Moreira, Santoro, Scudeller, Mesonero, Roade, Ruhlemann, Schaefer, Carrabba, Riveiro-Barciela, Basso, Valsecchi, Hernandez-Tejero, Acosta-Herrera, D’Angio, Baldini, Cazzaniga, Schulzky, Cecconi, Wittig, Ciccarelli, Rodriguez-Gandia, Bocciolone, Miozzo, Montano, Braun, Sacchi, Martinez, Ozer, Palmieri, Faverio, Preatoni, Bonfanti, Omodei, Tentorio, Castro, Rodrigues, Ortiz, de Cid, Ferrer, Gualtierotti, Nieto, Goerg, Badalamenti, Marsal, Matullo, Pelusi, Juzenas, Aliberti, Monzani, Moreno, Wesse, Lenz, Pumarola, Rimoldi, Bosari, Albrecht, Peter, Romero-Gomez, D’Amato, Duga, Banales, Hov, Folseraas, Valenti, Franke and Karlsen2020; COVID-19 Host Genetics Initiative, 2021; Pairo-Castineira et al., Reference Pairo-Castineira, Clohisey, Klaric, Bretherick, Rawlik, Pasko, Walker, Parkinson, Fourman, Russell, Furniss, Richmond, Gountouna, Wrobel, Harrison, Wang, Wu, Meynert, Griffiths, Oosthuyzen, Kousathanas, Moutsianas, Yang, Zhai, Zheng, Grimes, Beale, Millar, Shih, Keating, Zechner, Haley, Porteous, Hayward, Yang, Knight, Summers, Shankar-Hari, Klenerman, Turtle, Ho, Moore, Hinds, Horby, Nichol, Maslove, Ling, McAuley, Montgomery, Walsh, Pereira, Renieri, Gen, Investigators, Initiative, Me, Investigators, Gen, Shen, Ponting, Fawkes, Tenesa, Caulfield, Scott, Rowan, Murphy, PJM, Semple, Law, Vitart, Wilson and Baillie2021; Shelton et al., Reference Shelton, Shastri, Ye, Weldon, Filshtein-Sonmez, Coker, Symons, Esparza-Gordillo, Aslibekyan and Auton2021; Wu et al., Reference Wu, Ding, Li, Liu, Cheng, He, Xiao, Wu, Hou, Jiang, Long, Wang, Liu, Qu, Shi, Jiang, Mo, Ding, Fu, Han, Huo, Zeng, Zhou, Zhang, Ke, Xu, Ni, Shao, Wang, Liu, Li, Jin, Zheng, Wang, Liu, Li, Liu, Peng, Xu, Lin, Gao, Shi, Geng, Mu, Yan, Wang, Wu, Hao, Cheng, Qiu, Guo, Li, Chen, Sun, Lin, Jin, Wang, Sun and Wang2021; COVID-19 Host Genetics Initiative, 2022; Cruz et al., Reference Cruz, Almeida, Heredia, Quintela, Ceballos, Pita, Lorenzo-Salazar, Gonzalez-Montelongo, Gago-Dominguez, Porras, Castano, Nevado, Aguado, Aguilar, Aguilera-Albesa, Almadana, Almoguera, Alvarez, Andreu-Bernabeu, Arana-Arri, Arango, Arranz, Artiga, Baptista-Rosas, Barreda-Sanchez, Belhassen-Garcia, Bezerra, Bezerra, Boix-Palop, Brion, Brugada, Bustos, Calderon, Carbonell, Castano, Castelao, Conde-Vicente, Cordero-Lorenzana, Cortes-Sanchez, Corton, Darnaude, De Martino-Rodriguez, Campo-Perez, Bustamante, Dominguez-Garrido, Luchessi, Eiros, Sanabria, Farinas, Fernandez-Robelo, Fernandez-Rodriguez, Fernandez-Villa, Gil-Fournier, Gomez-Arrue, Alvarez, Quiros, Gonzalez-Penas, Gutierrez-Bautista, Herrero, Herrero-Gonzalez, Jimenez-Sousa, Lattig, Borja, Lopez-Rodriguez, Mancebo, Martin-Lopez, Martin, Martinez-Nieto, Martinez-Lopez, Martinez-Resendez, Martinez-Perez, Mazzeu, Macias, Minguez, Cuerda, Silbiger, Oliveira, Ortega-Paino, Parellada, Paz-Artal, Santos, Perez-Matute, Perez, Perez-Tomas, Perucho, Pinsach-Abuin, Pompa-Mera, Porras-Hurtado, Pujol, Leon, Resino, Fernandes, Rodriguez-Ruiz, Rodriguez-Artalejo, Rodriguez-Garcia, Ruiz-Cabello, Ruiz-Hornillos, Ryan, Soria, Souto, Tamayo, Tamayo-Velasco, Taracido-Fernandez, Teper, Torres-Tobar, Urioste, Valencia-Ramos, Yanez, Zarate, Nakanishi, Pigazzini, Degenhardt, Butler-Laporte, Maya-Miles, Bujanda, Bouysran, Palom, Ellinghaus, Martinez-Bueno, Rolker, Amitrano, Roade, Fava, Spinner, Prati, Bernardo, Garcia, Darcis, Fernandez-Cadenas, Holter, Banales, Frithiof, Duga, Asselta, Pereira, Romero-Gomez, Nafria-Jimenez, Hov, Migeotte, Renieri, Planas, Ludwig, Buti, Rahmouni, Alarcon-Riquelme, Schulte, Franke, Karlsen, Valenti, Zeberg, Richards, Ganna, Boada, Rojas, Ruiz, Sanchez, Real, Guillen-Navarro, Ayuso, Gonzalez-Neira, Riancho, Rojas-Martinez, Flores, Lapunzina and Carracedo2022; Degenhardt et al., Reference Degenhardt, Ellinghaus, Juzenas, Lerga-Jaso, Wendorff, Maya-Miles, Uellendahl-Werth, ElAbd, Ruhlemann, Arora, Ozer, Lenning, Myhre, Vadla, Wacker, Wienbrandt, Ortiz, Salazar, Chercoles, Palom, Ruiz, Garcia-Fernandez, Blanco-Grau, Mantovani, Zanella, Holten, Mayer, Bandera, Cherubini, Protti, Aghemo, Gerussi, Ramirez, Braun, Nebel, Barreira, Lleo, Teles, Kildal, Biondi, Caballero-Garralda, Ganna, Gori, Gluck, Lind, Tanck, Hinney, Nolla, Fracanzani, Peschuck, Cavallero, Dyrhol-Riise, Ruello, Julia, Muscatello, Pesenti, Voza, Rando-Segura, Solier, Schmidt, Cortes, Mateos, Nafria-Jimenez, Schaefer, Jensen, Bellinghausen, Maj, Ferrando, Horra, Quereda, Skurk, Thibeault, Scollo, Herr, Spinner, Gassner, Lange, Hu, Paccapelo, Lehmann, Angelini, Cappadona, Azuure, Bianco, Cea, Sancho, Hoff, Galimberti, Prati, Haschka, Jimenez, Pestana, Toapanta, Muniz-Diaz, Azzolini, Sandoval, Binatti, Scarpini, Helbig, Casalone, Urrechaga, Paraboschi, Pontali, Reverter, Calderon, Navas, Solligard, Contro, Arana-Arri, Aziz, Garcia, Sanchez, Ceriotti, Martinelli-Boneschi, Peyvandi, Kurth, Blasi, Malvestiti, Medrano, Mesonero, Rodriguez-Frias, Hanses, Muller, Hemmrich-Stanisak, Bellani, Grasselli, Pezzoli, Costantino, Albano, Cardamone, Bellelli, Citerio, Foti, Lamorte, Matullo, Baselli, Kurihara, Neb, My, Kurth, Hernandez, Pink, Rojas, Galvan-Femenia, Holter, Afset, Heyckendorf, Kassens, Damas, Rybniker, Altmuller, Ampuero, Martin, Erdmann, Banales, Badia, Dopazo, Schneider, Bergan, Barretina, Walter, Quero, Goikoetxea, Delgado, Guerrero, Fazaal, Kraft, Schroder, Risnes, Banasik, Muller, Gaede, Garcia-Etxebarria, Tonby, Heggelund, Izquierdo-Sanchez, Bettini, Sumoy, Sander, Lippert, Terranova, Nkambule, Knopp, Gustad, Garbarino, Santoro, Tellez, Roade, Ostadreza, Intxausti, Kogevinas, Riveiro-Barciela, Berger, Schaefer, Niemi, Gutierrez-Stampa, Carrabba, Figuera Basso, Valsecchi, Hernandez-Tejero, Vehreschild, Manunta, Acosta-Herrera, D’Angio, Baldini, Cazzaniga, Grimsrud, Cornberg, Nothen, Marquie, Castoldi, Cordioli, Cecconi, D’Amato, Augustin, Tomasi, Boada, Dreher, Seilmaier, Joannidis, Wittig, Mazzocco, Ciccarelli, Rodriguez-Gandia, Bocciolone, Miozzo, Ayo, Blay, Chueca, Montano, Braun, Ludwig, Marx, Martinez, Cornely, Witzke, Palmieri, Pa Study, Faverio, Preatoni, Bonfanti, Omodei, Tentorio, Castro, Rodrigues, Espana, Hoffmann, Rosenstiel, Schommers, Suwalski, Pablo, Ferrer, Bals, Gualtierotti, Gallego-Duran, Nieto, Carpani, Morilla, Badalamenti, Haider, Ciesek, May, Bombace, Marsal, Pigazzini, Klein, Pelusi, Wilfling, Bosari, Volland, Brunak, Raychaudhuri, Schreiber, Heilmann-Heimbach, Aliberti, Ripke, Dudman, Wesse, Zheng, Bahmer, Eggermann, Illig, Brenner, Pumarola, Feldt, Folseraas, Cejudo, Landmesser, Protzer, Hehr, Rimoldi, Monzani, Skogen, Keitel, Kopfnagel, Friaza, Andrade, Moreno, Albrecht, Peter, Poller, Farre, Yi, Wang, Khodamoradi, Karadeniz, Latiano, Goerg, Bacher, Koehler, Tran, Zoller, Schulte, Heidecker, Ludwig, Fernandez, Romero-Gomez, Albillos, Invernizzi, Buti, Duga, Bujanda, Hov, Lenz, Asselta, Cid, Valenti, Karlsen, Caceres and Franke2022; Horowitz et al., Reference Horowitz, Kosmicki, Damask, Sharma, Roberts, Justice, Banerjee, Coignet, Yadav, Leader, Marcketta, Park, Lanche, Maxwell, Knight, Bai, Guturu, Sun, Baltzell, Kury, Backman, Girshick, O’Dushlaine, McCurdy, Partha, Mansfield, Turissini, Li, Zhang, Mbatchou, Watanabe, Gurski, McCarthy, Kang, Dobbyn, Stahl, Verma, Sirugo, Genetics, Ritchie, Jones, Balasubramanian, Siminovitch, Salerno, Shuldiner, Rader, Mirshahi, Locke, Marchini, Overton, Carey, Habegger, Cantor, Rand, Hong, Reid, Ball, Baras, Abecasis and Ferreira2022; Namkoong et al., Reference Namkoong, Edahiro, Takano, Nishihara, Shirai, Sonehara, Tanaka, Azekawa, Mikami, Lee, Hasegawa, Okudela, Okuzaki, Motooka, Kanai, Naito, Yamamoto, Wang, Saiki, Ishihara, Matsubara, Hamamoto, Hayashi, Yoshimura, Tachikawa, Yanagita, Hyugaji, Shimizu, Katayama, Kato, Morita, Takahashi, Harada, Naito, Hiki, Matsushita, Takagi, Aoki, Nakamura, Harada, Sasano, Kabata, Masaki, Kamata, Ikemura, Chubachi, Okamori, Terai, Morita, Asakura, Sasaki, Morisaki, Uwamino, Nanki, Uchida, Uno, Nishimura, Ishiguro, Isono, Shibata, Matsui, Hosoda, Takano, Nishida, Kobayashi, Takaku, Takayanagi, Ueda, Tada, Miyawaki, Yamamoto, Yoshida, Hayashi, Nagasaka, Arai, Kaneko, Sasaki, Tagaya, Kawana, Arimura, Takahashi, Anzai, Ito, Endo, Uchimura, Miyazaki, Honda, Tateishi, Tohda, Ichimura, Sonobe, Sassa, Nakajima, Nakano, Nakajima, Anan, Arai, Kurihara, Harada, Nishio, Ueda, Azuma, Saito, Sado, Miyazaki, Sato, Haruta, Nagasaki, Yasui, Hasegawa, Mutoh, Kimura, Sato, Takei, Hagimoto, Noguchi, Yamano, Sasano, Ota, Nakamori, Yoshiya, Saito, Yoshihara, Wada, Iwamura, Kanayama, Maruyama, Yoshiyama, Ohta, Kokuto, Ogata, Tanaka, Arakawa, Shimoda, Osawa, Tateno, Hase, Yoshida, Suzuki, Kawada, Horinouchi, Saito, Mitamura, Hagihara, Ochi, Uchida, Baba, Arai, Ogura, Takahashi, Hagiwara, Nagao, Konishi, Nakachi, Murakami, Yamada, Sugiura, Sano, Matsumoto, Kimura, Ono, Baba, Suzuki, Nakayama, Masuzawa, Namba, Suzuki, Naito, Liu, Takuwa, Sugihara, Wing, Sakakibara, Hizawa, Shiroyama, Miyawaki, Kawamura, Nakayama, Matsuo, Maeda, Nii, Noda, Niitsu, Adachi, Enomoto, Amiya, Hara, Yamaguchi, Murakami, Kuge, Matsumoto, Yamamoto, Yamamoto, Yoneda, Kishikawa, Yamada, Kawabata, Kijima, Takagaki, Sasa, Ueno, Suzuki, Takemoto, Eguchi, Fukusumi, Imai, Fukushima, Kishima, Inohara, Tomono, Kato, Takahashi, Matsuda, Hirata, Takeda, Koh, Manabe, Funatsu, Ito, Fukui, Shinozuka, Kohashi, Miyazaki, Shoko, Kojima, Adachi, Ishikawa, Takahashi, Inoue, Hirano, Kobayashi, Takaoka, Watanabe, Miyazawa, Kimura, Sado, Sugimoto, Kamiya, Kuwahara, Fujiwara, Matsunaga, Sato, Okada, Hirai, Kawashima, Narita, Niwa, Sekikawa, Nishi, Nishitsuji, Tani, Suzuki, Nakatsumi, Ogura, Kitamura, Hagiwara, Murohashi, Okabayashi, Mochimaru, Nukaga, Satomi, Oyamada, Mori, Baba, Fukui, Odate, Mashimo, Makino, Yagi, Hashiguchi, Kagyo, Shiomi, Fuke, Saito, Tsuchida, Fujitani, Takita, Morikawa, Yoshida, Izumo, Inomata, Kuse, Awano, Tone, Ito, Nakamura, Hoshino, Maruyama, Ishikura, Takata, Odani, Amishima, Hattori, Shichinohe, Kagaya, Kita, Ohta, Sakagami, Koshida, Hayashi, Shimizu, Kozu, Hiranuma, Gon, Izumi, Nagata, Ueda, Taki, Hanada, Kawamura, Ichikado, Nishiyama, Muranaka, Nakamura, Hashimoto, Wakahara, Koji, Omote, Ando, Kodama, Kaneyama, Maeda, Kuraki, Matsumoto, Yokote, Nakada, Abe, Oshima, Shimada, Harada, Takahashi, Ono, Sakurai, Shibusawa, Kimizuka, Kawana, Sano, Watanabe, Suematsu, Sageshima, Yoshifuji, Ito, Takahashi, Ishioka, Nakamura, Masuda, Wakabayashi, Watanabe, Ueda, Nishikawa, Chihara, Takeuchi, Onoi, Shinozuka, Sueyoshi, Nagasaki, Okamoto, Ishihara, Shimo, Tokunaga, Kusaka, Ohba, Isogai, Ogawa, Inoue, Fukuyama, Eriguchi, Yonekawa, Kan, Matsumoto, Kanaoka, Ihara, Komuta, Inoue, Chiba, Yamagata, Hiramatsu, Kai, Asano, Oguma, Ito, Hashimoto, Yamasaki, Kasamatsu, Komase, Hida, Tsuburai, Oyama, Takada, Kanda, Kitagawa, Fukuta, Miyake, Yoshida, Ogura, Abe, Kono, Togashi, Takoi, Kikuchi, Ogawa, Ogata, Ishihara, Kanehiro, Ozaki, Fuchimoto, Wada, Fujimoto, Nishiyama, Terashima, Beppu, Yoshida, Narumoto, Nagai, Ooshima, Motegi, Umeda, Miyagawa, Shimada, Endo, Ohira, Watanabe, Inoue, Igarashi, Sato, Sagara, Tanaka, Ohta, Kimura, Shibata, Tanino, Nikaido, Minemura, Sato, Yamada, Hashino, Shinoki, Iwagoe, Takahashi, Fujii, Kishi, Kanai, Imamura, Yamashita, Yatomi, Maeno, Hayashi, Takahashi, Kuramochi, Kamimaki, Tominaga, Ishii, Utsugi, Ono, Tanaka, Kashiwada, Fujita, Saito, Seike, Watanabe, Matsuse, Kodaka, Nakano, Oshio, Hirouchi, Makino, Egi, Omae, Nannya, Ueno, Katayama, Ai, Fukui, Kumanogoh, Sato, Hasegawa, Tokunaga, Ishii, Koike, Kitagawa, Kimura, Imoto, Miyano, Ogawa, Kanai, Fukunaga and Okada2022; Roberts et al., Reference Roberts, Partha, Rhead, Knight, Park, Coignet, Zhang, Berkowitz, Turrisini, Gaddis, McCurdy, Pavlovic, Ruiz, Sass, Ancestry, Baltzell, Guturu, Girshick, Ball, Hong and Rand2022; Table 1), locus 9q34.2 with the ABO gene stands out because the genetic association there points directly to the ABO gene and because the risk/protective effect inferred from the ABO blood group distribution among cases and controls is very similar between genome-wide and blood group candidate studies. The genetic variants at the ABO gene locus represent the statistically strongest genetic associations in the so-called Manhattan P value association plots of the GWAS studies (Shelton et al., Reference Shelton, Shastri, Ye, Weldon, Filshtein-Sonmez, Coker, Symons, Esparza-Gordillo, Aslibekyan and Auton2021) (or sometimes the second strongest next to the association signal at locus 3p21.31 (Ellinghaus et al., Reference Ellinghaus, Degenhardt, Bujanda, Buti, Albillos, Invernizzi, Fernandez, Prati, Baselli, Asselta, Grimsrud, Milani, Aziz, Kassens, May, Wendorff, Wienbrandt, Uellendahl-Werth, Zheng, Yi, de Pablo, Chercoles, Palom, Garcia-Fernandez, Rodriguez-Frias, Zanella, Bandera, Protti, Aghemo, Lleo, Biondi, Caballero-Garralda, Gori, Tanck, Carreras Nolla, Latiano, Fracanzani, Peschuck, Julia, Pesenti, Voza, Jimenez, Mateos, Jimenez, Quereda, Paccapelo, Gassner, Angelini, Cea, Solier, Pestana, Muniz-Diaz, Sandoval, Paraboschi, Navas, Garcia Sanchez, Ceriotti, Martinelli-Boneschi, Peyvandi, Blasi, Tellez, Blanco-Grau, Hemmrich-Stanisak, Grasselli, Costantino, Cardamone, Foti, Aneli, Kurihara, ElAbd, My, Galvan-Femenia, Martin, Erdmann, Ferrusquia-Acosta, Garcia-Etxebarria, Izquierdo-Sanchez, Bettini, Sumoy, Terranova, Moreira, Santoro, Scudeller, Mesonero, Roade, Ruhlemann, Schaefer, Carrabba, Riveiro-Barciela, Basso, Valsecchi, Hernandez-Tejero, Acosta-Herrera, D’Angio, Baldini, Cazzaniga, Schulzky, Cecconi, Wittig, Ciccarelli, Rodriguez-Gandia, Bocciolone, Miozzo, Montano, Braun, Sacchi, Martinez, Ozer, Palmieri, Faverio, Preatoni, Bonfanti, Omodei, Tentorio, Castro, Rodrigues, Ortiz, de Cid, Ferrer, Gualtierotti, Nieto, Goerg, Badalamenti, Marsal, Matullo, Pelusi, Juzenas, Aliberti, Monzani, Moreno, Wesse, Lenz, Pumarola, Rimoldi, Bosari, Albrecht, Peter, Romero-Gomez, D’Amato, Duga, Banales, Hov, Folseraas, Valenti, Franke and Karlsen2020; Horowitz et al., Reference Horowitz, Kosmicki, Damask, Sharma, Roberts, Justice, Banerjee, Coignet, Yadav, Leader, Marcketta, Park, Lanche, Maxwell, Knight, Bai, Guturu, Sun, Baltzell, Kury, Backman, Girshick, O’Dushlaine, McCurdy, Partha, Mansfield, Turissini, Li, Zhang, Mbatchou, Watanabe, Gurski, McCarthy, Kang, Dobbyn, Stahl, Verma, Sirugo, Genetics, Ritchie, Jones, Balasubramanian, Siminovitch, Salerno, Shuldiner, Rader, Mirshahi, Locke, Marchini, Overton, Carey, Habegger, Cantor, Rand, Hong, Reid, Ball, Baras, Abecasis and Ferreira2022), depending on whether one is testing for disease severity or infection). Moreover, it was shown that the genetic association at the ABO locus cannot be explained by COVID-19 comorbidities, that is, potential confounding factors (Ellinghaus et al., Reference Ellinghaus, Degenhardt, Bujanda, Buti, Albillos, Invernizzi, Fernandez, Prati, Baselli, Asselta, Grimsrud, Milani, Aziz, Kassens, May, Wendorff, Wienbrandt, Uellendahl-Werth, Zheng, Yi, de Pablo, Chercoles, Palom, Garcia-Fernandez, Rodriguez-Frias, Zanella, Bandera, Protti, Aghemo, Lleo, Biondi, Caballero-Garralda, Gori, Tanck, Carreras Nolla, Latiano, Fracanzani, Peschuck, Julia, Pesenti, Voza, Jimenez, Mateos, Jimenez, Quereda, Paccapelo, Gassner, Angelini, Cea, Solier, Pestana, Muniz-Diaz, Sandoval, Paraboschi, Navas, Garcia Sanchez, Ceriotti, Martinelli-Boneschi, Peyvandi, Blasi, Tellez, Blanco-Grau, Hemmrich-Stanisak, Grasselli, Costantino, Cardamone, Foti, Aneli, Kurihara, ElAbd, My, Galvan-Femenia, Martin, Erdmann, Ferrusquia-Acosta, Garcia-Etxebarria, Izquierdo-Sanchez, Bettini, Sumoy, Terranova, Moreira, Santoro, Scudeller, Mesonero, Roade, Ruhlemann, Schaefer, Carrabba, Riveiro-Barciela, Basso, Valsecchi, Hernandez-Tejero, Acosta-Herrera, D’Angio, Baldini, Cazzaniga, Schulzky, Cecconi, Wittig, Ciccarelli, Rodriguez-Gandia, Bocciolone, Miozzo, Montano, Braun, Sacchi, Martinez, Ozer, Palmieri, Faverio, Preatoni, Bonfanti, Omodei, Tentorio, Castro, Rodrigues, Ortiz, de Cid, Ferrer, Gualtierotti, Nieto, Goerg, Badalamenti, Marsal, Matullo, Pelusi, Juzenas, Aliberti, Monzani, Moreno, Wesse, Lenz, Pumarola, Rimoldi, Bosari, Albrecht, Peter, Romero-Gomez, D’Amato, Duga, Banales, Hov, Folseraas, Valenti, Franke and Karlsen2020; Horowitz et al., Reference Horowitz, Kosmicki, Damask, Sharma, Roberts, Justice, Banerjee, Coignet, Yadav, Leader, Marcketta, Park, Lanche, Maxwell, Knight, Bai, Guturu, Sun, Baltzell, Kury, Backman, Girshick, O’Dushlaine, McCurdy, Partha, Mansfield, Turissini, Li, Zhang, Mbatchou, Watanabe, Gurski, McCarthy, Kang, Dobbyn, Stahl, Verma, Sirugo, Genetics, Ritchie, Jones, Balasubramanian, Siminovitch, Salerno, Shuldiner, Rader, Mirshahi, Locke, Marchini, Overton, Carey, Habegger, Cantor, Rand, Hong, Reid, Ball, Baras, Abecasis and Ferreira2022). It should be noted that most candidate genes listed in Table 1 from the respective publications, with the exception of ABO and ACE2 (here a rare variant association upstream of the angiotensin-converting enzyme 2 gene, the primary cell entry receptor for SARS-CoV-2, was identified (Horowitz et al., Reference Horowitz, Kosmicki, Damask, Sharma, Roberts, Justice, Banerjee, Coignet, Yadav, Leader, Marcketta, Park, Lanche, Maxwell, Knight, Bai, Guturu, Sun, Baltzell, Kury, Backman, Girshick, O’Dushlaine, McCurdy, Partha, Mansfield, Turissini, Li, Zhang, Mbatchou, Watanabe, Gurski, McCarthy, Kang, Dobbyn, Stahl, Verma, Sirugo, Genetics, Ritchie, Jones, Balasubramanian, Siminovitch, Salerno, Shuldiner, Rader, Mirshahi, Locke, Marchini, Overton, Carey, Habegger, Cantor, Rand, Hong, Reid, Ball, Baras, Abecasis and Ferreira2022) which, however, describes a much smaller proportion of the heritability (So et al., Reference So, Gui, Cherny and Sham2011) for COVID-19 susceptibility compared to the ABO association in the general population due to its rare frequency and its comparable effect size with those of the common variants), are so far predominantly candidate genes that need to be investigated in functional studies for a biological effect. Indeed, most of the loci listed in Table 1 span a large number of genes, so that many genes at a susceptibility locus may be candidate susceptibility genes. An important observation about the ABO locus is that the ABO association signal has been replicated in almost all large COVID-19 GWAS studies, making it the most replicated locus for COVID-19 (along with 3p21.31). Multiple genetic variants have been identified for the ABO locus (Ellinghaus et al., Reference Ellinghaus, Degenhardt, Bujanda, Buti, Albillos, Invernizzi, Fernandez, Prati, Baselli, Asselta, Grimsrud, Milani, Aziz, Kassens, May, Wendorff, Wienbrandt, Uellendahl-Werth, Zheng, Yi, de Pablo, Chercoles, Palom, Garcia-Fernandez, Rodriguez-Frias, Zanella, Bandera, Protti, Aghemo, Lleo, Biondi, Caballero-Garralda, Gori, Tanck, Carreras Nolla, Latiano, Fracanzani, Peschuck, Julia, Pesenti, Voza, Jimenez, Mateos, Jimenez, Quereda, Paccapelo, Gassner, Angelini, Cea, Solier, Pestana, Muniz-Diaz, Sandoval, Paraboschi, Navas, Garcia Sanchez, Ceriotti, Martinelli-Boneschi, Peyvandi, Blasi, Tellez, Blanco-Grau, Hemmrich-Stanisak, Grasselli, Costantino, Cardamone, Foti, Aneli, Kurihara, ElAbd, My, Galvan-Femenia, Martin, Erdmann, Ferrusquia-Acosta, Garcia-Etxebarria, Izquierdo-Sanchez, Bettini, Sumoy, Terranova, Moreira, Santoro, Scudeller, Mesonero, Roade, Ruhlemann, Schaefer, Carrabba, Riveiro-Barciela, Basso, Valsecchi, Hernandez-Tejero, Acosta-Herrera, D’Angio, Baldini, Cazzaniga, Schulzky, Cecconi, Wittig, Ciccarelli, Rodriguez-Gandia, Bocciolone, Miozzo, Montano, Braun, Sacchi, Martinez, Ozer, Palmieri, Faverio, Preatoni, Bonfanti, Omodei, Tentorio, Castro, Rodrigues, Ortiz, de Cid, Ferrer, Gualtierotti, Nieto, Goerg, Badalamenti, Marsal, Matullo, Pelusi, Juzenas, Aliberti, Monzani, Moreno, Wesse, Lenz, Pumarola, Rimoldi, Bosari, Albrecht, Peter, Romero-Gomez, D’Amato, Duga, Banales, Hov, Folseraas, Valenti, Franke and Karlsen2020; COVID-19 Host Genetics Initiative, 2021; Shelton et al., Reference Shelton, Shastri, Ye, Weldon, Filshtein-Sonmez, Coker, Symons, Esparza-Gordillo, Aslibekyan and Auton2021; Cruz et al., Reference Cruz, Almeida, Heredia, Quintela, Ceballos, Pita, Lorenzo-Salazar, Gonzalez-Montelongo, Gago-Dominguez, Porras, Castano, Nevado, Aguado, Aguilar, Aguilera-Albesa, Almadana, Almoguera, Alvarez, Andreu-Bernabeu, Arana-Arri, Arango, Arranz, Artiga, Baptista-Rosas, Barreda-Sanchez, Belhassen-Garcia, Bezerra, Bezerra, Boix-Palop, Brion, Brugada, Bustos, Calderon, Carbonell, Castano, Castelao, Conde-Vicente, Cordero-Lorenzana, Cortes-Sanchez, Corton, Darnaude, De Martino-Rodriguez, Campo-Perez, Bustamante, Dominguez-Garrido, Luchessi, Eiros, Sanabria, Farinas, Fernandez-Robelo, Fernandez-Rodriguez, Fernandez-Villa, Gil-Fournier, Gomez-Arrue, Alvarez, Quiros, Gonzalez-Penas, Gutierrez-Bautista, Herrero, Herrero-Gonzalez, Jimenez-Sousa, Lattig, Borja, Lopez-Rodriguez, Mancebo, Martin-Lopez, Martin, Martinez-Nieto, Martinez-Lopez, Martinez-Resendez, Martinez-Perez, Mazzeu, Macias, Minguez, Cuerda, Silbiger, Oliveira, Ortega-Paino, Parellada, Paz-Artal, Santos, Perez-Matute, Perez, Perez-Tomas, Perucho, Pinsach-Abuin, Pompa-Mera, Porras-Hurtado, Pujol, Leon, Resino, Fernandes, Rodriguez-Ruiz, Rodriguez-Artalejo, Rodriguez-Garcia, Ruiz-Cabello, Ruiz-Hornillos, Ryan, Soria, Souto, Tamayo, Tamayo-Velasco, Taracido-Fernandez, Teper, Torres-Tobar, Urioste, Valencia-Ramos, Yanez, Zarate, Nakanishi, Pigazzini, Degenhardt, Butler-Laporte, Maya-Miles, Bujanda, Bouysran, Palom, Ellinghaus, Martinez-Bueno, Rolker, Amitrano, Roade, Fava, Spinner, Prati, Bernardo, Garcia, Darcis, Fernandez-Cadenas, Holter, Banales, Frithiof, Duga, Asselta, Pereira, Romero-Gomez, Nafria-Jimenez, Hov, Migeotte, Renieri, Planas, Ludwig, Buti, Rahmouni, Alarcon-Riquelme, Schulte, Franke, Karlsen, Valenti, Zeberg, Richards, Ganna, Boada, Rojas, Ruiz, Sanchez, Real, Guillen-Navarro, Ayuso, Gonzalez-Neira, Riancho, Rojas-Martinez, Flores, Lapunzina and Carracedo2022; Degenhardt et al., Reference Degenhardt, Ellinghaus, Juzenas, Lerga-Jaso, Wendorff, Maya-Miles, Uellendahl-Werth, ElAbd, Ruhlemann, Arora, Ozer, Lenning, Myhre, Vadla, Wacker, Wienbrandt, Ortiz, Salazar, Chercoles, Palom, Ruiz, Garcia-Fernandez, Blanco-Grau, Mantovani, Zanella, Holten, Mayer, Bandera, Cherubini, Protti, Aghemo, Gerussi, Ramirez, Braun, Nebel, Barreira, Lleo, Teles, Kildal, Biondi, Caballero-Garralda, Ganna, Gori, Gluck, Lind, Tanck, Hinney, Nolla, Fracanzani, Peschuck, Cavallero, Dyrhol-Riise, Ruello, Julia, Muscatello, Pesenti, Voza, Rando-Segura, Solier, Schmidt, Cortes, Mateos, Nafria-Jimenez, Schaefer, Jensen, Bellinghausen, Maj, Ferrando, Horra, Quereda, Skurk, Thibeault, Scollo, Herr, Spinner, Gassner, Lange, Hu, Paccapelo, Lehmann, Angelini, Cappadona, Azuure, Bianco, Cea, Sancho, Hoff, Galimberti, Prati, Haschka, Jimenez, Pestana, Toapanta, Muniz-Diaz, Azzolini, Sandoval, Binatti, Scarpini, Helbig, Casalone, Urrechaga, Paraboschi, Pontali, Reverter, Calderon, Navas, Solligard, Contro, Arana-Arri, Aziz, Garcia, Sanchez, Ceriotti, Martinelli-Boneschi, Peyvandi, Kurth, Blasi, Malvestiti, Medrano, Mesonero, Rodriguez-Frias, Hanses, Muller, Hemmrich-Stanisak, Bellani, Grasselli, Pezzoli, Costantino, Albano, Cardamone, Bellelli, Citerio, Foti, Lamorte, Matullo, Baselli, Kurihara, Neb, My, Kurth, Hernandez, Pink, Rojas, Galvan-Femenia, Holter, Afset, Heyckendorf, Kassens, Damas, Rybniker, Altmuller, Ampuero, Martin, Erdmann, Banales, Badia, Dopazo, Schneider, Bergan, Barretina, Walter, Quero, Goikoetxea, Delgado, Guerrero, Fazaal, Kraft, Schroder, Risnes, Banasik, Muller, Gaede, Garcia-Etxebarria, Tonby, Heggelund, Izquierdo-Sanchez, Bettini, Sumoy, Sander, Lippert, Terranova, Nkambule, Knopp, Gustad, Garbarino, Santoro, Tellez, Roade, Ostadreza, Intxausti, Kogevinas, Riveiro-Barciela, Berger, Schaefer, Niemi, Gutierrez-Stampa, Carrabba, Figuera Basso, Valsecchi, Hernandez-Tejero, Vehreschild, Manunta, Acosta-Herrera, D’Angio, Baldini, Cazzaniga, Grimsrud, Cornberg, Nothen, Marquie, Castoldi, Cordioli, Cecconi, D’Amato, Augustin, Tomasi, Boada, Dreher, Seilmaier, Joannidis, Wittig, Mazzocco, Ciccarelli, Rodriguez-Gandia, Bocciolone, Miozzo, Ayo, Blay, Chueca, Montano, Braun, Ludwig, Marx, Martinez, Cornely, Witzke, Palmieri, Pa Study, Faverio, Preatoni, Bonfanti, Omodei, Tentorio, Castro, Rodrigues, Espana, Hoffmann, Rosenstiel, Schommers, Suwalski, Pablo, Ferrer, Bals, Gualtierotti, Gallego-Duran, Nieto, Carpani, Morilla, Badalamenti, Haider, Ciesek, May, Bombace, Marsal, Pigazzini, Klein, Pelusi, Wilfling, Bosari, Volland, Brunak, Raychaudhuri, Schreiber, Heilmann-Heimbach, Aliberti, Ripke, Dudman, Wesse, Zheng, Bahmer, Eggermann, Illig, Brenner, Pumarola, Feldt, Folseraas, Cejudo, Landmesser, Protzer, Hehr, Rimoldi, Monzani, Skogen, Keitel, Kopfnagel, Friaza, Andrade, Moreno, Albrecht, Peter, Poller, Farre, Yi, Wang, Khodamoradi, Karadeniz, Latiano, Goerg, Bacher, Koehler, Tran, Zoller, Schulte, Heidecker, Ludwig, Fernandez, Romero-Gomez, Albillos, Invernizzi, Buti, Duga, Bujanda, Hov, Lenz, Asselta, Cid, Valenti, Karlsen, Caceres and Franke2022; Horowitz et al., Reference Horowitz, Kosmicki, Damask, Sharma, Roberts, Justice, Banerjee, Coignet, Yadav, Leader, Marcketta, Park, Lanche, Maxwell, Knight, Bai, Guturu, Sun, Baltzell, Kury, Backman, Girshick, O’Dushlaine, McCurdy, Partha, Mansfield, Turissini, Li, Zhang, Mbatchou, Watanabe, Gurski, McCarthy, Kang, Dobbyn, Stahl, Verma, Sirugo, Genetics, Ritchie, Jones, Balasubramanian, Siminovitch, Salerno, Shuldiner, Rader, Mirshahi, Locke, Marchini, Overton, Carey, Habegger, Cantor, Rand, Hong, Reid, Ball, Baras, Abecasis and Ferreira2022; Roberts et al., Reference Roberts, Partha, Rhead, Knight, Park, Coignet, Zhang, Berkowitz, Turrisini, Gaddis, McCurdy, Pavlovic, Ruiz, Sass, Ancestry, Baltzell, Guturu, Girshick, Ball, Hong and Rand2022; Table 2), including a frameshift insertion (rs8176719) in a recent trans-ethnic GWAS meta-analysis (Wu et al., Reference Wu, Ding, Li, Liu, Cheng, He, Xiao, Wu, Hou, Jiang, Long, Wang, Liu, Qu, Shi, Jiang, Mo, Ding, Fu, Han, Huo, Zeng, Zhou, Zhang, Ke, Xu, Ni, Shao, Wang, Liu, Li, Jin, Zheng, Wang, Liu, Li, Liu, Peng, Xu, Lin, Gao, Shi, Geng, Mu, Yan, Wang, Wu, Hao, Cheng, Qiu, Guo, Li, Chen, Sun, Lin, Jin, Wang, Sun and Wang2021), although in this genome-wide meta-analysis, the frameshift polymorphism was only identified in one GWAS study from China using sequencing data and further confirmation is needed. The same is true for the association of a genetic variant near the DOCK2 gene, which so far is significant only in one GWAS study from Japan and for the age group <65 years (Namkoong et al., Reference Namkoong, Edahiro, Takano, Nishihara, Shirai, Sonehara, Tanaka, Azekawa, Mikami, Lee, Hasegawa, Okudela, Okuzaki, Motooka, Kanai, Naito, Yamamoto, Wang, Saiki, Ishihara, Matsubara, Hamamoto, Hayashi, Yoshimura, Tachikawa, Yanagita, Hyugaji, Shimizu, Katayama, Kato, Morita, Takahashi, Harada, Naito, Hiki, Matsushita, Takagi, Aoki, Nakamura, Harada, Sasano, Kabata, Masaki, Kamata, Ikemura, Chubachi, Okamori, Terai, Morita, Asakura, Sasaki, Morisaki, Uwamino, Nanki, Uchida, Uno, Nishimura, Ishiguro, Isono, Shibata, Matsui, Hosoda, Takano, Nishida, Kobayashi, Takaku, Takayanagi, Ueda, Tada, Miyawaki, Yamamoto, Yoshida, Hayashi, Nagasaka, Arai, Kaneko, Sasaki, Tagaya, Kawana, Arimura, Takahashi, Anzai, Ito, Endo, Uchimura, Miyazaki, Honda, Tateishi, Tohda, Ichimura, Sonobe, Sassa, Nakajima, Nakano, Nakajima, Anan, Arai, Kurihara, Harada, Nishio, Ueda, Azuma, Saito, Sado, Miyazaki, Sato, Haruta, Nagasaki, Yasui, Hasegawa, Mutoh, Kimura, Sato, Takei, Hagimoto, Noguchi, Yamano, Sasano, Ota, Nakamori, Yoshiya, Saito, Yoshihara, Wada, Iwamura, Kanayama, Maruyama, Yoshiyama, Ohta, Kokuto, Ogata, Tanaka, Arakawa, Shimoda, Osawa, Tateno, Hase, Yoshida, Suzuki, Kawada, Horinouchi, Saito, Mitamura, Hagihara, Ochi, Uchida, Baba, Arai, Ogura, Takahashi, Hagiwara, Nagao, Konishi, Nakachi, Murakami, Yamada, Sugiura, Sano, Matsumoto, Kimura, Ono, Baba, Suzuki, Nakayama, Masuzawa, Namba, Suzuki, Naito, Liu, Takuwa, Sugihara, Wing, Sakakibara, Hizawa, Shiroyama, Miyawaki, Kawamura, Nakayama, Matsuo, Maeda, Nii, Noda, Niitsu, Adachi, Enomoto, Amiya, Hara, Yamaguchi, Murakami, Kuge, Matsumoto, Yamamoto, Yamamoto, Yoneda, Kishikawa, Yamada, Kawabata, Kijima, Takagaki, Sasa, Ueno, Suzuki, Takemoto, Eguchi, Fukusumi, Imai, Fukushima, Kishima, Inohara, Tomono, Kato, Takahashi, Matsuda, Hirata, Takeda, Koh, Manabe, Funatsu, Ito, Fukui, Shinozuka, Kohashi, Miyazaki, Shoko, Kojima, Adachi, Ishikawa, Takahashi, Inoue, Hirano, Kobayashi, Takaoka, Watanabe, Miyazawa, Kimura, Sado, Sugimoto, Kamiya, Kuwahara, Fujiwara, Matsunaga, Sato, Okada, Hirai, Kawashima, Narita, Niwa, Sekikawa, Nishi, Nishitsuji, Tani, Suzuki, Nakatsumi, Ogura, Kitamura, Hagiwara, Murohashi, Okabayashi, Mochimaru, Nukaga, Satomi, Oyamada, Mori, Baba, Fukui, Odate, Mashimo, Makino, Yagi, Hashiguchi, Kagyo, Shiomi, Fuke, Saito, Tsuchida, Fujitani, Takita, Morikawa, Yoshida, Izumo, Inomata, Kuse, Awano, Tone, Ito, Nakamura, Hoshino, Maruyama, Ishikura, Takata, Odani, Amishima, Hattori, Shichinohe, Kagaya, Kita, Ohta, Sakagami, Koshida, Hayashi, Shimizu, Kozu, Hiranuma, Gon, Izumi, Nagata, Ueda, Taki, Hanada, Kawamura, Ichikado, Nishiyama, Muranaka, Nakamura, Hashimoto, Wakahara, Koji, Omote, Ando, Kodama, Kaneyama, Maeda, Kuraki, Matsumoto, Yokote, Nakada, Abe, Oshima, Shimada, Harada, Takahashi, Ono, Sakurai, Shibusawa, Kimizuka, Kawana, Sano, Watanabe, Suematsu, Sageshima, Yoshifuji, Ito, Takahashi, Ishioka, Nakamura, Masuda, Wakabayashi, Watanabe, Ueda, Nishikawa, Chihara, Takeuchi, Onoi, Shinozuka, Sueyoshi, Nagasaki, Okamoto, Ishihara, Shimo, Tokunaga, Kusaka, Ohba, Isogai, Ogawa, Inoue, Fukuyama, Eriguchi, Yonekawa, Kan, Matsumoto, Kanaoka, Ihara, Komuta, Inoue, Chiba, Yamagata, Hiramatsu, Kai, Asano, Oguma, Ito, Hashimoto, Yamasaki, Kasamatsu, Komase, Hida, Tsuburai, Oyama, Takada, Kanda, Kitagawa, Fukuta, Miyake, Yoshida, Ogura, Abe, Kono, Togashi, Takoi, Kikuchi, Ogawa, Ogata, Ishihara, Kanehiro, Ozaki, Fuchimoto, Wada, Fujimoto, Nishiyama, Terashima, Beppu, Yoshida, Narumoto, Nagai, Ooshima, Motegi, Umeda, Miyagawa, Shimada, Endo, Ohira, Watanabe, Inoue, Igarashi, Sato, Sagara, Tanaka, Ohta, Kimura, Shibata, Tanino, Nikaido, Minemura, Sato, Yamada, Hashino, Shinoki, Iwagoe, Takahashi, Fujii, Kishi, Kanai, Imamura, Yamashita, Yatomi, Maeno, Hayashi, Takahashi, Kuramochi, Kamimaki, Tominaga, Ishii, Utsugi, Ono, Tanaka, Kashiwada, Fujita, Saito, Seike, Watanabe, Matsuse, Kodaka, Nakano, Oshio, Hirouchi, Makino, Egi, Omae, Nannya, Ueno, Katayama, Ai, Fukui, Kumanogoh, Sato, Hasegawa, Tokunaga, Ishii, Koike, Kitagawa, Kimura, Imoto, Miyano, Ogawa, Kanai, Fukunaga and Okada2022).
Table 1. Genome-wide significant (P < 5 × 10−8) susceptibility loci for SARS-CoV-2 infection and/or COVID-19 disease severity identified in large-scale (hypothesis-free) genome-wide analyses to date (as of August 2022) (Ellinghaus et al., Reference Ellinghaus, Degenhardt, Bujanda, Buti, Albillos, Invernizzi, Fernandez, Prati, Baselli, Asselta, Grimsrud, Milani, Aziz, Kassens, May, Wendorff, Wienbrandt, Uellendahl-Werth, Zheng, Yi, de Pablo, Chercoles, Palom, Garcia-Fernandez, Rodriguez-Frias, Zanella, Bandera, Protti, Aghemo, Lleo, Biondi, Caballero-Garralda, Gori, Tanck, Carreras Nolla, Latiano, Fracanzani, Peschuck, Julia, Pesenti, Voza, Jimenez, Mateos, Jimenez, Quereda, Paccapelo, Gassner, Angelini, Cea, Solier, Pestana, Muniz-Diaz, Sandoval, Paraboschi, Navas, Garcia Sanchez, Ceriotti, Martinelli-Boneschi, Peyvandi, Blasi, Tellez, Blanco-Grau, Hemmrich-Stanisak, Grasselli, Costantino, Cardamone, Foti, Aneli, Kurihara, ElAbd, My, Galvan-Femenia, Martin, Erdmann, Ferrusquia-Acosta, Garcia-Etxebarria, Izquierdo-Sanchez, Bettini, Sumoy, Terranova, Moreira, Santoro, Scudeller, Mesonero, Roade, Ruhlemann, Schaefer, Carrabba, Riveiro-Barciela, Basso, Valsecchi, Hernandez-Tejero, Acosta-Herrera, D’Angio, Baldini, Cazzaniga, Schulzky, Cecconi, Wittig, Ciccarelli, Rodriguez-Gandia, Bocciolone, Miozzo, Montano, Braun, Sacchi, Martinez, Ozer, Palmieri, Faverio, Preatoni, Bonfanti, Omodei, Tentorio, Castro, Rodrigues, Ortiz, de Cid, Ferrer, Gualtierotti, Nieto, Goerg, Badalamenti, Marsal, Matullo, Pelusi, Juzenas, Aliberti, Monzani, Moreno, Wesse, Lenz, Pumarola, Rimoldi, Bosari, Albrecht, Peter, Romero-Gomez, D’Amato, Duga, Banales, Hov, Folseraas, Valenti, Franke and Karlsen2020; COVID-19 Host Genetics Initiative, 2021; Pairo-Castineira et al., Reference Pairo-Castineira, Clohisey, Klaric, Bretherick, Rawlik, Pasko, Walker, Parkinson, Fourman, Russell, Furniss, Richmond, Gountouna, Wrobel, Harrison, Wang, Wu, Meynert, Griffiths, Oosthuyzen, Kousathanas, Moutsianas, Yang, Zhai, Zheng, Grimes, Beale, Millar, Shih, Keating, Zechner, Haley, Porteous, Hayward, Yang, Knight, Summers, Shankar-Hari, Klenerman, Turtle, Ho, Moore, Hinds, Horby, Nichol, Maslove, Ling, McAuley, Montgomery, Walsh, Pereira, Renieri, Gen, Investigators, Initiative, Me, Investigators, Gen, Shen, Ponting, Fawkes, Tenesa, Caulfield, Scott, Rowan, Murphy, PJM, Semple, Law, Vitart, Wilson and Baillie2021; Shelton et al., Reference Shelton, Shastri, Ye, Weldon, Filshtein-Sonmez, Coker, Symons, Esparza-Gordillo, Aslibekyan and Auton2021; Wu et al., Reference Wu, Ding, Li, Liu, Cheng, He, Xiao, Wu, Hou, Jiang, Long, Wang, Liu, Qu, Shi, Jiang, Mo, Ding, Fu, Han, Huo, Zeng, Zhou, Zhang, Ke, Xu, Ni, Shao, Wang, Liu, Li, Jin, Zheng, Wang, Liu, Li, Liu, Peng, Xu, Lin, Gao, Shi, Geng, Mu, Yan, Wang, Wu, Hao, Cheng, Qiu, Guo, Li, Chen, Sun, Lin, Jin, Wang, Sun and Wang2021; COVID-19 Host Genetics Initiative, 2022; Cruz et al., Reference Cruz, Almeida, Heredia, Quintela, Ceballos, Pita, Lorenzo-Salazar, Gonzalez-Montelongo, Gago-Dominguez, Porras, Castano, Nevado, Aguado, Aguilar, Aguilera-Albesa, Almadana, Almoguera, Alvarez, Andreu-Bernabeu, Arana-Arri, Arango, Arranz, Artiga, Baptista-Rosas, Barreda-Sanchez, Belhassen-Garcia, Bezerra, Bezerra, Boix-Palop, Brion, Brugada, Bustos, Calderon, Carbonell, Castano, Castelao, Conde-Vicente, Cordero-Lorenzana, Cortes-Sanchez, Corton, Darnaude, De Martino-Rodriguez, Campo-Perez, Bustamante, Dominguez-Garrido, Luchessi, Eiros, Sanabria, Farinas, Fernandez-Robelo, Fernandez-Rodriguez, Fernandez-Villa, Gil-Fournier, Gomez-Arrue, Alvarez, Quiros, Gonzalez-Penas, Gutierrez-Bautista, Herrero, Herrero-Gonzalez, Jimenez-Sousa, Lattig, Borja, Lopez-Rodriguez, Mancebo, Martin-Lopez, Martin, Martinez-Nieto, Martinez-Lopez, Martinez-Resendez, Martinez-Perez, Mazzeu, Macias, Minguez, Cuerda, Silbiger, Oliveira, Ortega-Paino, Parellada, Paz-Artal, Santos, Perez-Matute, Perez, Perez-Tomas, Perucho, Pinsach-Abuin, Pompa-Mera, Porras-Hurtado, Pujol, Leon, Resino, Fernandes, Rodriguez-Ruiz, Rodriguez-Artalejo, Rodriguez-Garcia, Ruiz-Cabello, Ruiz-Hornillos, Ryan, Soria, Souto, Tamayo, Tamayo-Velasco, Taracido-Fernandez, Teper, Torres-Tobar, Urioste, Valencia-Ramos, Yanez, Zarate, Nakanishi, Pigazzini, Degenhardt, Butler-Laporte, Maya-Miles, Bujanda, Bouysran, Palom, Ellinghaus, Martinez-Bueno, Rolker, Amitrano, Roade, Fava, Spinner, Prati, Bernardo, Garcia, Darcis, Fernandez-Cadenas, Holter, Banales, Frithiof, Duga, Asselta, Pereira, Romero-Gomez, Nafria-Jimenez, Hov, Migeotte, Renieri, Planas, Ludwig, Buti, Rahmouni, Alarcon-Riquelme, Schulte, Franke, Karlsen, Valenti, Zeberg, Richards, Ganna, Boada, Rojas, Ruiz, Sanchez, Real, Guillen-Navarro, Ayuso, Gonzalez-Neira, Riancho, Rojas-Martinez, Flores, Lapunzina and Carracedo2022; Degenhardt et al., Reference Degenhardt, Ellinghaus, Juzenas, Lerga-Jaso, Wendorff, Maya-Miles, Uellendahl-Werth, ElAbd, Ruhlemann, Arora, Ozer, Lenning, Myhre, Vadla, Wacker, Wienbrandt, Ortiz, Salazar, Chercoles, Palom, Ruiz, Garcia-Fernandez, Blanco-Grau, Mantovani, Zanella, Holten, Mayer, Bandera, Cherubini, Protti, Aghemo, Gerussi, Ramirez, Braun, Nebel, Barreira, Lleo, Teles, Kildal, Biondi, Caballero-Garralda, Ganna, Gori, Gluck, Lind, Tanck, Hinney, Nolla, Fracanzani, Peschuck, Cavallero, Dyrhol-Riise, Ruello, Julia, Muscatello, Pesenti, Voza, Rando-Segura, Solier, Schmidt, Cortes, Mateos, Nafria-Jimenez, Schaefer, Jensen, Bellinghausen, Maj, Ferrando, Horra, Quereda, Skurk, Thibeault, Scollo, Herr, Spinner, Gassner, Lange, Hu, Paccapelo, Lehmann, Angelini, Cappadona, Azuure, Bianco, Cea, Sancho, Hoff, Galimberti, Prati, Haschka, Jimenez, Pestana, Toapanta, Muniz-Diaz, Azzolini, Sandoval, Binatti, Scarpini, Helbig, Casalone, Urrechaga, Paraboschi, Pontali, Reverter, Calderon, Navas, Solligard, Contro, Arana-Arri, Aziz, Garcia, Sanchez, Ceriotti, Martinelli-Boneschi, Peyvandi, Kurth, Blasi, Malvestiti, Medrano, Mesonero, Rodriguez-Frias, Hanses, Muller, Hemmrich-Stanisak, Bellani, Grasselli, Pezzoli, Costantino, Albano, Cardamone, Bellelli, Citerio, Foti, Lamorte, Matullo, Baselli, Kurihara, Neb, My, Kurth, Hernandez, Pink, Rojas, Galvan-Femenia, Holter, Afset, Heyckendorf, Kassens, Damas, Rybniker, Altmuller, Ampuero, Martin, Erdmann, Banales, Badia, Dopazo, Schneider, Bergan, Barretina, Walter, Quero, Goikoetxea, Delgado, Guerrero, Fazaal, Kraft, Schroder, Risnes, Banasik, Muller, Gaede, Garcia-Etxebarria, Tonby, Heggelund, Izquierdo-Sanchez, Bettini, Sumoy, Sander, Lippert, Terranova, Nkambule, Knopp, Gustad, Garbarino, Santoro, Tellez, Roade, Ostadreza, Intxausti, Kogevinas, Riveiro-Barciela, Berger, Schaefer, Niemi, Gutierrez-Stampa, Carrabba, Figuera Basso, Valsecchi, Hernandez-Tejero, Vehreschild, Manunta, Acosta-Herrera, D’Angio, Baldini, Cazzaniga, Grimsrud, Cornberg, Nothen, Marquie, Castoldi, Cordioli, Cecconi, D’Amato, Augustin, Tomasi, Boada, Dreher, Seilmaier, Joannidis, Wittig, Mazzocco, Ciccarelli, Rodriguez-Gandia, Bocciolone, Miozzo, Ayo, Blay, Chueca, Montano, Braun, Ludwig, Marx, Martinez, Cornely, Witzke, Palmieri, Pa Study, Faverio, Preatoni, Bonfanti, Omodei, Tentorio, Castro, Rodrigues, Espana, Hoffmann, Rosenstiel, Schommers, Suwalski, Pablo, Ferrer, Bals, Gualtierotti, Gallego-Duran, Nieto, Carpani, Morilla, Badalamenti, Haider, Ciesek, May, Bombace, Marsal, Pigazzini, Klein, Pelusi, Wilfling, Bosari, Volland, Brunak, Raychaudhuri, Schreiber, Heilmann-Heimbach, Aliberti, Ripke, Dudman, Wesse, Zheng, Bahmer, Eggermann, Illig, Brenner, Pumarola, Feldt, Folseraas, Cejudo, Landmesser, Protzer, Hehr, Rimoldi, Monzani, Skogen, Keitel, Kopfnagel, Friaza, Andrade, Moreno, Albrecht, Peter, Poller, Farre, Yi, Wang, Khodamoradi, Karadeniz, Latiano, Goerg, Bacher, Koehler, Tran, Zoller, Schulte, Heidecker, Ludwig, Fernandez, Romero-Gomez, Albillos, Invernizzi, Buti, Duga, Bujanda, Hov, Lenz, Asselta, Cid, Valenti, Karlsen, Caceres and Franke2022; Horowitz et al., Reference Horowitz, Kosmicki, Damask, Sharma, Roberts, Justice, Banerjee, Coignet, Yadav, Leader, Marcketta, Park, Lanche, Maxwell, Knight, Bai, Guturu, Sun, Baltzell, Kury, Backman, Girshick, O’Dushlaine, McCurdy, Partha, Mansfield, Turissini, Li, Zhang, Mbatchou, Watanabe, Gurski, McCarthy, Kang, Dobbyn, Stahl, Verma, Sirugo, Genetics, Ritchie, Jones, Balasubramanian, Siminovitch, Salerno, Shuldiner, Rader, Mirshahi, Locke, Marchini, Overton, Carey, Habegger, Cantor, Rand, Hong, Reid, Ball, Baras, Abecasis and Ferreira2022; Namkoong et al., Reference Namkoong, Edahiro, Takano, Nishihara, Shirai, Sonehara, Tanaka, Azekawa, Mikami, Lee, Hasegawa, Okudela, Okuzaki, Motooka, Kanai, Naito, Yamamoto, Wang, Saiki, Ishihara, Matsubara, Hamamoto, Hayashi, Yoshimura, Tachikawa, Yanagita, Hyugaji, Shimizu, Katayama, Kato, Morita, Takahashi, Harada, Naito, Hiki, Matsushita, Takagi, Aoki, Nakamura, Harada, Sasano, Kabata, Masaki, Kamata, Ikemura, Chubachi, Okamori, Terai, Morita, Asakura, Sasaki, Morisaki, Uwamino, Nanki, Uchida, Uno, Nishimura, Ishiguro, Isono, Shibata, Matsui, Hosoda, Takano, Nishida, Kobayashi, Takaku, Takayanagi, Ueda, Tada, Miyawaki, Yamamoto, Yoshida, Hayashi, Nagasaka, Arai, Kaneko, Sasaki, Tagaya, Kawana, Arimura, Takahashi, Anzai, Ito, Endo, Uchimura, Miyazaki, Honda, Tateishi, Tohda, Ichimura, Sonobe, Sassa, Nakajima, Nakano, Nakajima, Anan, Arai, Kurihara, Harada, Nishio, Ueda, Azuma, Saito, Sado, Miyazaki, Sato, Haruta, Nagasaki, Yasui, Hasegawa, Mutoh, Kimura, Sato, Takei, Hagimoto, Noguchi, Yamano, Sasano, Ota, Nakamori, Yoshiya, Saito, Yoshihara, Wada, Iwamura, Kanayama, Maruyama, Yoshiyama, Ohta, Kokuto, Ogata, Tanaka, Arakawa, Shimoda, Osawa, Tateno, Hase, Yoshida, Suzuki, Kawada, Horinouchi, Saito, Mitamura, Hagihara, Ochi, Uchida, Baba, Arai, Ogura, Takahashi, Hagiwara, Nagao, Konishi, Nakachi, Murakami, Yamada, Sugiura, Sano, Matsumoto, Kimura, Ono, Baba, Suzuki, Nakayama, Masuzawa, Namba, Suzuki, Naito, Liu, Takuwa, Sugihara, Wing, Sakakibara, Hizawa, Shiroyama, Miyawaki, Kawamura, Nakayama, Matsuo, Maeda, Nii, Noda, Niitsu, Adachi, Enomoto, Amiya, Hara, Yamaguchi, Murakami, Kuge, Matsumoto, Yamamoto, Yamamoto, Yoneda, Kishikawa, Yamada, Kawabata, Kijima, Takagaki, Sasa, Ueno, Suzuki, Takemoto, Eguchi, Fukusumi, Imai, Fukushima, Kishima, Inohara, Tomono, Kato, Takahashi, Matsuda, Hirata, Takeda, Koh, Manabe, Funatsu, Ito, Fukui, Shinozuka, Kohashi, Miyazaki, Shoko, Kojima, Adachi, Ishikawa, Takahashi, Inoue, Hirano, Kobayashi, Takaoka, Watanabe, Miyazawa, Kimura, Sado, Sugimoto, Kamiya, Kuwahara, Fujiwara, Matsunaga, Sato, Okada, Hirai, Kawashima, Narita, Niwa, Sekikawa, Nishi, Nishitsuji, Tani, Suzuki, Nakatsumi, Ogura, Kitamura, Hagiwara, Murohashi, Okabayashi, Mochimaru, Nukaga, Satomi, Oyamada, Mori, Baba, Fukui, Odate, Mashimo, Makino, Yagi, Hashiguchi, Kagyo, Shiomi, Fuke, Saito, Tsuchida, Fujitani, Takita, Morikawa, Yoshida, Izumo, Inomata, Kuse, Awano, Tone, Ito, Nakamura, Hoshino, Maruyama, Ishikura, Takata, Odani, Amishima, Hattori, Shichinohe, Kagaya, Kita, Ohta, Sakagami, Koshida, Hayashi, Shimizu, Kozu, Hiranuma, Gon, Izumi, Nagata, Ueda, Taki, Hanada, Kawamura, Ichikado, Nishiyama, Muranaka, Nakamura, Hashimoto, Wakahara, Koji, Omote, Ando, Kodama, Kaneyama, Maeda, Kuraki, Matsumoto, Yokote, Nakada, Abe, Oshima, Shimada, Harada, Takahashi, Ono, Sakurai, Shibusawa, Kimizuka, Kawana, Sano, Watanabe, Suematsu, Sageshima, Yoshifuji, Ito, Takahashi, Ishioka, Nakamura, Masuda, Wakabayashi, Watanabe, Ueda, Nishikawa, Chihara, Takeuchi, Onoi, Shinozuka, Sueyoshi, Nagasaki, Okamoto, Ishihara, Shimo, Tokunaga, Kusaka, Ohba, Isogai, Ogawa, Inoue, Fukuyama, Eriguchi, Yonekawa, Kan, Matsumoto, Kanaoka, Ihara, Komuta, Inoue, Chiba, Yamagata, Hiramatsu, Kai, Asano, Oguma, Ito, Hashimoto, Yamasaki, Kasamatsu, Komase, Hida, Tsuburai, Oyama, Takada, Kanda, Kitagawa, Fukuta, Miyake, Yoshida, Ogura, Abe, Kono, Togashi, Takoi, Kikuchi, Ogawa, Ogata, Ishihara, Kanehiro, Ozaki, Fuchimoto, Wada, Fujimoto, Nishiyama, Terashima, Beppu, Yoshida, Narumoto, Nagai, Ooshima, Motegi, Umeda, Miyagawa, Shimada, Endo, Ohira, Watanabe, Inoue, Igarashi, Sato, Sagara, Tanaka, Ohta, Kimura, Shibata, Tanino, Nikaido, Minemura, Sato, Yamada, Hashino, Shinoki, Iwagoe, Takahashi, Fujii, Kishi, Kanai, Imamura, Yamashita, Yatomi, Maeno, Hayashi, Takahashi, Kuramochi, Kamimaki, Tominaga, Ishii, Utsugi, Ono, Tanaka, Kashiwada, Fujita, Saito, Seike, Watanabe, Matsuse, Kodaka, Nakano, Oshio, Hirouchi, Makino, Egi, Omae, Nannya, Ueno, Katayama, Ai, Fukui, Kumanogoh, Sato, Hasegawa, Tokunaga, Ishii, Koike, Kitagawa, Kimura, Imoto, Miyano, Ogawa, Kanai, Fukunaga and Okada2022; Roberts et al., Reference Roberts, Partha, Rhead, Knight, Park, Coignet, Zhang, Berkowitz, Turrisini, Gaddis, McCurdy, Pavlovic, Ruiz, Sass, Ancestry, Baltzell, Guturu, Girshick, Ball, Hong and Rand2022)

Note: Susceptibility variants from GWAS and GWMA studies with fewer than 1,500 individuals and fewer than two study populations as well as candidate gene studies are not listed. Variant: dbSNP name of the lead variant (without mentioning nearby variant names from later studies). Locus: chromosomal region. Chromosome: position: position according to human genome build hg38. EA: effect allele. NEA: non-effect allele. OR: odds ratio (i.e., estimated effect size) with respect to EA from the respective publication. The effect direction refers to whether the EA increases (OR > 1) or decreases (OR < 1) the risk of infection and/or disease severity. Candidate gene: preferably selected candidate gene from the respective publication. Reference: publication in which the genome-wide significant association was first reported.
Table 2. Genome-wide significant associations between genetic variants at the ABO gene locus and SARS-CoV-2 infection and/or COVID-19 disease severity (including other important phenotypic associations for the variants)

The ABO blood group system in brief
The ABO histo-blood group system includes two antigens (A and B) and four blood groups (A, B, AB and O). The ABO blood group antigens, which are expressed predominantly on N-linked and O-linked glycoproteins as well as glycolipids, are expressed not only on erythrocytes but also on numerous other cell types. Their synthesis first requires synthesis of the histo-blood group H precursor antigen, which is catalyzed by the enzymes (fucosyltransferases) FUT1 (e.g., in erythroblasts, megakaryocytes and vascular endothelial cells) or FUT2 (in epithelial cells of, e.g., the upper respiratory tract and digestive tract), and then blood group A or B enzymes (glycosyltransferases) generate the A and B antigens (Cooling, Reference Cooling2015). In many epithelial tissues, ABO expression is highly dependent on the inheritance of the Secretor/FUT2 gene, and null alleles of FUT2 (the “nonsecretor” phenotype) are very common in the population (approximately 5–50% worldwide (Nordgren et al., Reference Nordgren, Sharma, Kambhampati, Lopman and Svensson2016)), resulting in a deficiency of precursor H antigen synthesis and thus also a deficiency of A and B antigens in the corresponding cell types. The ABO gene and the FUT2 gene are two of the few human genes that are clearly subject to frequency-dependent balanced selection (Pendu et al., Reference Pendu, Breiman, Rocher, Dion and Ruvoen-Clouet2021), which also suggests an important role in their interaction with environmental factors such as gut microbes (Ruhlemann et al., Reference Ruhlemann, Hermes, Bang, Doms, Moitinho-Silva, Thingholm, Frost, Degenhardt, Wittig, Kassens, Weiss, Peters, Neuhaus, Volker, Volzke, Homuth, Weiss, Grallert, Laudes, Lieb, Haller, Lerch, Baines and Franke2021). The glycosyltransferases of blood groups A and B are encoded by different alleles of the ABO gene: Type A antigen is synthesized by the glycosyltransferase encoded by A alleles of the ABO gene, while type B antigen is synthesized by the glycosyltransferase encoded by B alleles. The A and B antigens are autosomal codominant; this means that both A and B antigens are synthesized in A/B individuals. Blood group O is autosomal recessive, and the O alleles are unable to produce a functional enzyme; therefore, in O/O individuals, the H precursor antigen is left unchanged. For this reason, O blood group individuals have anti-A and anti-B antibodies, A blood group individuals have anti-B antibodies, B blood group individuals have anti-A antibodies and AB blood group individuals have neither anti-A nor anti-B antibodies.
Are the ABO locus and ABO blood groups related to infection risk, disease severity or both in COVID-19?
Although initial genome-wide and candidate studies have not yet provided a clear picture of this question, it is now apparent that genetic variants at the ABO locus confer risk (or protection) with SARS-CoV-2 infection and COVID-19 severity (Figure 1 and Table 2). Numerous hypothesis-driven (nongenome-wide) studies have also reported associations between ABO blood groups and risk for COVID-19 infection (Barnkob et al., Reference Barnkob, Pottegard, Stovring, Haunstrup, Homburg, Larsen, Hansen, Titlestad, Aagaard, Moller and Barington2020; Goker et al., Reference Goker, Karakulak, Demiroglu, Ceylan, Buyukasik, Inkaya, Aksu, Sayinalp, Haznedaroglu, Uzun, Akova, Ozcebe and Unal2020; Leaf et al., Reference Leaf, Al-Samkari, Brenner, Gupta and Leaf2020; Li et al., Reference Li, Wang, Chen, Cai, Deng and Yang2020; Zietz et al., Reference Zietz, Zucker and Tatonetti2020; Ahmed et al., Reference Ahmed, Quinn and Tan2021; Solmaz and Arac, Reference Solmaz and Arac2021; Zhao et al., Reference Zhao, Yang, Huang, Li, Gu, Lu, Zhang, Liu, Liu, Liu, He, Sun, Wei, Yang, Wang, Zhang, Zhou, Xing and Wang2021) and severity of COVID-19 (Hoiland et al., Reference Hoiland, Fergusson, Mitra, Griesdale, Devine, Stukas, Cooper, Thiara, Foster, Chen, Lee, Conway, Wellington and Sekhon2020; Sardu et al., Reference Sardu, Marfella, Maggi, Messina, Cirillo, Codella, Gambardella, Sardu, Gatta, Santulli and Paolisso2020; Muniz-Diaz et al., Reference Muniz-Diaz, Llopis, Parra, Roig, Ferrer, Grifols, Millan, Ene, Ramiro, Maglio, Garcia, Pinacho, Jaramillo, Pero, Artaza, Valles, Sauleda, Puig and Contreras2021; Ray et al., Reference Ray, Schull, Vermeulen and Park2021) with almost all studies reaching the same conclusion as the GWAS/GWMA studies: A lower risk of infection for people with blood type O than for people with non-O blood types, with blood type A (sometimes AB (Namkoong et al., Reference Namkoong, Edahiro, Takano, Nishihara, Shirai, Sonehara, Tanaka, Azekawa, Mikami, Lee, Hasegawa, Okudela, Okuzaki, Motooka, Kanai, Naito, Yamamoto, Wang, Saiki, Ishihara, Matsubara, Hamamoto, Hayashi, Yoshimura, Tachikawa, Yanagita, Hyugaji, Shimizu, Katayama, Kato, Morita, Takahashi, Harada, Naito, Hiki, Matsushita, Takagi, Aoki, Nakamura, Harada, Sasano, Kabata, Masaki, Kamata, Ikemura, Chubachi, Okamori, Terai, Morita, Asakura, Sasaki, Morisaki, Uwamino, Nanki, Uchida, Uno, Nishimura, Ishiguro, Isono, Shibata, Matsui, Hosoda, Takano, Nishida, Kobayashi, Takaku, Takayanagi, Ueda, Tada, Miyawaki, Yamamoto, Yoshida, Hayashi, Nagasaka, Arai, Kaneko, Sasaki, Tagaya, Kawana, Arimura, Takahashi, Anzai, Ito, Endo, Uchimura, Miyazaki, Honda, Tateishi, Tohda, Ichimura, Sonobe, Sassa, Nakajima, Nakano, Nakajima, Anan, Arai, Kurihara, Harada, Nishio, Ueda, Azuma, Saito, Sado, Miyazaki, Sato, Haruta, Nagasaki, Yasui, Hasegawa, Mutoh, Kimura, Sato, Takei, Hagimoto, Noguchi, Yamano, Sasano, Ota, Nakamori, Yoshiya, Saito, Yoshihara, Wada, Iwamura, Kanayama, Maruyama, Yoshiyama, Ohta, Kokuto, Ogata, Tanaka, Arakawa, Shimoda, Osawa, Tateno, Hase, Yoshida, Suzuki, Kawada, Horinouchi, Saito, Mitamura, Hagihara, Ochi, Uchida, Baba, Arai, Ogura, Takahashi, Hagiwara, Nagao, Konishi, Nakachi, Murakami, Yamada, Sugiura, Sano, Matsumoto, Kimura, Ono, Baba, Suzuki, Nakayama, Masuzawa, Namba, Suzuki, Naito, Liu, Takuwa, Sugihara, Wing, Sakakibara, Hizawa, Shiroyama, Miyawaki, Kawamura, Nakayama, Matsuo, Maeda, Nii, Noda, Niitsu, Adachi, Enomoto, Amiya, Hara, Yamaguchi, Murakami, Kuge, Matsumoto, Yamamoto, Yamamoto, Yoneda, Kishikawa, Yamada, Kawabata, Kijima, Takagaki, Sasa, Ueno, Suzuki, Takemoto, Eguchi, Fukusumi, Imai, Fukushima, Kishima, Inohara, Tomono, Kato, Takahashi, Matsuda, Hirata, Takeda, Koh, Manabe, Funatsu, Ito, Fukui, Shinozuka, Kohashi, Miyazaki, Shoko, Kojima, Adachi, Ishikawa, Takahashi, Inoue, Hirano, Kobayashi, Takaoka, Watanabe, Miyazawa, Kimura, Sado, Sugimoto, Kamiya, Kuwahara, Fujiwara, Matsunaga, Sato, Okada, Hirai, Kawashima, Narita, Niwa, Sekikawa, Nishi, Nishitsuji, Tani, Suzuki, Nakatsumi, Ogura, Kitamura, Hagiwara, Murohashi, Okabayashi, Mochimaru, Nukaga, Satomi, Oyamada, Mori, Baba, Fukui, Odate, Mashimo, Makino, Yagi, Hashiguchi, Kagyo, Shiomi, Fuke, Saito, Tsuchida, Fujitani, Takita, Morikawa, Yoshida, Izumo, Inomata, Kuse, Awano, Tone, Ito, Nakamura, Hoshino, Maruyama, Ishikura, Takata, Odani, Amishima, Hattori, Shichinohe, Kagaya, Kita, Ohta, Sakagami, Koshida, Hayashi, Shimizu, Kozu, Hiranuma, Gon, Izumi, Nagata, Ueda, Taki, Hanada, Kawamura, Ichikado, Nishiyama, Muranaka, Nakamura, Hashimoto, Wakahara, Koji, Omote, Ando, Kodama, Kaneyama, Maeda, Kuraki, Matsumoto, Yokote, Nakada, Abe, Oshima, Shimada, Harada, Takahashi, Ono, Sakurai, Shibusawa, Kimizuka, Kawana, Sano, Watanabe, Suematsu, Sageshima, Yoshifuji, Ito, Takahashi, Ishioka, Nakamura, Masuda, Wakabayashi, Watanabe, Ueda, Nishikawa, Chihara, Takeuchi, Onoi, Shinozuka, Sueyoshi, Nagasaki, Okamoto, Ishihara, Shimo, Tokunaga, Kusaka, Ohba, Isogai, Ogawa, Inoue, Fukuyama, Eriguchi, Yonekawa, Kan, Matsumoto, Kanaoka, Ihara, Komuta, Inoue, Chiba, Yamagata, Hiramatsu, Kai, Asano, Oguma, Ito, Hashimoto, Yamasaki, Kasamatsu, Komase, Hida, Tsuburai, Oyama, Takada, Kanda, Kitagawa, Fukuta, Miyake, Yoshida, Ogura, Abe, Kono, Togashi, Takoi, Kikuchi, Ogawa, Ogata, Ishihara, Kanehiro, Ozaki, Fuchimoto, Wada, Fujimoto, Nishiyama, Terashima, Beppu, Yoshida, Narumoto, Nagai, Ooshima, Motegi, Umeda, Miyagawa, Shimada, Endo, Ohira, Watanabe, Inoue, Igarashi, Sato, Sagara, Tanaka, Ohta, Kimura, Shibata, Tanino, Nikaido, Minemura, Sato, Yamada, Hashino, Shinoki, Iwagoe, Takahashi, Fujii, Kishi, Kanai, Imamura, Yamashita, Yatomi, Maeno, Hayashi, Takahashi, Kuramochi, Kamimaki, Tominaga, Ishii, Utsugi, Ono, Tanaka, Kashiwada, Fujita, Saito, Seike, Watanabe, Matsuse, Kodaka, Nakano, Oshio, Hirouchi, Makino, Egi, Omae, Nannya, Ueno, Katayama, Ai, Fukui, Kumanogoh, Sato, Hasegawa, Tokunaga, Ishii, Koike, Kitagawa, Kimura, Imoto, Miyano, Ogawa, Kanai, Fukunaga and Okada2022) because of higher number of patients examined in Asian countries where AB is more common) being associated with a higher risk. Although the significant estimated risk (odds ratio(OR) > 1) or protection (OR < 1) of the ABO blood groups is rather small (for SARS-CoV-2 infection: OR 0.81 and 95% confidence interval (95% CI) 0.75–0.86 for O vs. A/B/AB estimated across 20 cohort studies (Franchini et al., Reference Franchini, Cruciani, Mengoli, Marano, Candura, Lopez, Pati, Pupella and De Angelis2021); for COVID-19 disease severity with respiratory support: OR 0.65 for O vs. A/B/AB in Italian/Spanish study populations (Ellinghaus et al., Reference Ellinghaus, Degenhardt, Bujanda, Buti, Albillos, Invernizzi, Fernandez, Prati, Baselli, Asselta, Grimsrud, Milani, Aziz, Kassens, May, Wendorff, Wienbrandt, Uellendahl-Werth, Zheng, Yi, de Pablo, Chercoles, Palom, Garcia-Fernandez, Rodriguez-Frias, Zanella, Bandera, Protti, Aghemo, Lleo, Biondi, Caballero-Garralda, Gori, Tanck, Carreras Nolla, Latiano, Fracanzani, Peschuck, Julia, Pesenti, Voza, Jimenez, Mateos, Jimenez, Quereda, Paccapelo, Gassner, Angelini, Cea, Solier, Pestana, Muniz-Diaz, Sandoval, Paraboschi, Navas, Garcia Sanchez, Ceriotti, Martinelli-Boneschi, Peyvandi, Blasi, Tellez, Blanco-Grau, Hemmrich-Stanisak, Grasselli, Costantino, Cardamone, Foti, Aneli, Kurihara, ElAbd, My, Galvan-Femenia, Martin, Erdmann, Ferrusquia-Acosta, Garcia-Etxebarria, Izquierdo-Sanchez, Bettini, Sumoy, Terranova, Moreira, Santoro, Scudeller, Mesonero, Roade, Ruhlemann, Schaefer, Carrabba, Riveiro-Barciela, Basso, Valsecchi, Hernandez-Tejero, Acosta-Herrera, D’Angio, Baldini, Cazzaniga, Schulzky, Cecconi, Wittig, Ciccarelli, Rodriguez-Gandia, Bocciolone, Miozzo, Montano, Braun, Sacchi, Martinez, Ozer, Palmieri, Faverio, Preatoni, Bonfanti, Omodei, Tentorio, Castro, Rodrigues, Ortiz, de Cid, Ferrer, Gualtierotti, Nieto, Goerg, Badalamenti, Marsal, Matullo, Pelusi, Juzenas, Aliberti, Monzani, Moreno, Wesse, Lenz, Pumarola, Rimoldi, Bosari, Albrecht, Peter, Romero-Gomez, D’Amato, Duga, Banales, Hov, Folseraas, Valenti, Franke and Karlsen2020); OR 0.81 for O vs. A/B/AB for Japanese study populations (Namkoong et al., Reference Namkoong, Edahiro, Takano, Nishihara, Shirai, Sonehara, Tanaka, Azekawa, Mikami, Lee, Hasegawa, Okudela, Okuzaki, Motooka, Kanai, Naito, Yamamoto, Wang, Saiki, Ishihara, Matsubara, Hamamoto, Hayashi, Yoshimura, Tachikawa, Yanagita, Hyugaji, Shimizu, Katayama, Kato, Morita, Takahashi, Harada, Naito, Hiki, Matsushita, Takagi, Aoki, Nakamura, Harada, Sasano, Kabata, Masaki, Kamata, Ikemura, Chubachi, Okamori, Terai, Morita, Asakura, Sasaki, Morisaki, Uwamino, Nanki, Uchida, Uno, Nishimura, Ishiguro, Isono, Shibata, Matsui, Hosoda, Takano, Nishida, Kobayashi, Takaku, Takayanagi, Ueda, Tada, Miyawaki, Yamamoto, Yoshida, Hayashi, Nagasaka, Arai, Kaneko, Sasaki, Tagaya, Kawana, Arimura, Takahashi, Anzai, Ito, Endo, Uchimura, Miyazaki, Honda, Tateishi, Tohda, Ichimura, Sonobe, Sassa, Nakajima, Nakano, Nakajima, Anan, Arai, Kurihara, Harada, Nishio, Ueda, Azuma, Saito, Sado, Miyazaki, Sato, Haruta, Nagasaki, Yasui, Hasegawa, Mutoh, Kimura, Sato, Takei, Hagimoto, Noguchi, Yamano, Sasano, Ota, Nakamori, Yoshiya, Saito, Yoshihara, Wada, Iwamura, Kanayama, Maruyama, Yoshiyama, Ohta, Kokuto, Ogata, Tanaka, Arakawa, Shimoda, Osawa, Tateno, Hase, Yoshida, Suzuki, Kawada, Horinouchi, Saito, Mitamura, Hagihara, Ochi, Uchida, Baba, Arai, Ogura, Takahashi, Hagiwara, Nagao, Konishi, Nakachi, Murakami, Yamada, Sugiura, Sano, Matsumoto, Kimura, Ono, Baba, Suzuki, Nakayama, Masuzawa, Namba, Suzuki, Naito, Liu, Takuwa, Sugihara, Wing, Sakakibara, Hizawa, Shiroyama, Miyawaki, Kawamura, Nakayama, Matsuo, Maeda, Nii, Noda, Niitsu, Adachi, Enomoto, Amiya, Hara, Yamaguchi, Murakami, Kuge, Matsumoto, Yamamoto, Yamamoto, Yoneda, Kishikawa, Yamada, Kawabata, Kijima, Takagaki, Sasa, Ueno, Suzuki, Takemoto, Eguchi, Fukusumi, Imai, Fukushima, Kishima, Inohara, Tomono, Kato, Takahashi, Matsuda, Hirata, Takeda, Koh, Manabe, Funatsu, Ito, Fukui, Shinozuka, Kohashi, Miyazaki, Shoko, Kojima, Adachi, Ishikawa, Takahashi, Inoue, Hirano, Kobayashi, Takaoka, Watanabe, Miyazawa, Kimura, Sado, Sugimoto, Kamiya, Kuwahara, Fujiwara, Matsunaga, Sato, Okada, Hirai, Kawashima, Narita, Niwa, Sekikawa, Nishi, Nishitsuji, Tani, Suzuki, Nakatsumi, Ogura, Kitamura, Hagiwara, Murohashi, Okabayashi, Mochimaru, Nukaga, Satomi, Oyamada, Mori, Baba, Fukui, Odate, Mashimo, Makino, Yagi, Hashiguchi, Kagyo, Shiomi, Fuke, Saito, Tsuchida, Fujitani, Takita, Morikawa, Yoshida, Izumo, Inomata, Kuse, Awano, Tone, Ito, Nakamura, Hoshino, Maruyama, Ishikura, Takata, Odani, Amishima, Hattori, Shichinohe, Kagaya, Kita, Ohta, Sakagami, Koshida, Hayashi, Shimizu, Kozu, Hiranuma, Gon, Izumi, Nagata, Ueda, Taki, Hanada, Kawamura, Ichikado, Nishiyama, Muranaka, Nakamura, Hashimoto, Wakahara, Koji, Omote, Ando, Kodama, Kaneyama, Maeda, Kuraki, Matsumoto, Yokote, Nakada, Abe, Oshima, Shimada, Harada, Takahashi, Ono, Sakurai, Shibusawa, Kimizuka, Kawana, Sano, Watanabe, Suematsu, Sageshima, Yoshifuji, Ito, Takahashi, Ishioka, Nakamura, Masuda, Wakabayashi, Watanabe, Ueda, Nishikawa, Chihara, Takeuchi, Onoi, Shinozuka, Sueyoshi, Nagasaki, Okamoto, Ishihara, Shimo, Tokunaga, Kusaka, Ohba, Isogai, Ogawa, Inoue, Fukuyama, Eriguchi, Yonekawa, Kan, Matsumoto, Kanaoka, Ihara, Komuta, Inoue, Chiba, Yamagata, Hiramatsu, Kai, Asano, Oguma, Ito, Hashimoto, Yamasaki, Kasamatsu, Komase, Hida, Tsuburai, Oyama, Takada, Kanda, Kitagawa, Fukuta, Miyake, Yoshida, Ogura, Abe, Kono, Togashi, Takoi, Kikuchi, Ogawa, Ogata, Ishihara, Kanehiro, Ozaki, Fuchimoto, Wada, Fujimoto, Nishiyama, Terashima, Beppu, Yoshida, Narumoto, Nagai, Ooshima, Motegi, Umeda, Miyagawa, Shimada, Endo, Ohira, Watanabe, Inoue, Igarashi, Sato, Sagara, Tanaka, Ohta, Kimura, Shibata, Tanino, Nikaido, Minemura, Sato, Yamada, Hashino, Shinoki, Iwagoe, Takahashi, Fujii, Kishi, Kanai, Imamura, Yamashita, Yatomi, Maeno, Hayashi, Takahashi, Kuramochi, Kamimaki, Tominaga, Ishii, Utsugi, Ono, Tanaka, Kashiwada, Fujita, Saito, Seike, Watanabe, Matsuse, Kodaka, Nakano, Oshio, Hirouchi, Makino, Egi, Omae, Nannya, Ueno, Katayama, Ai, Fukui, Kumanogoh, Sato, Hasegawa, Tokunaga, Ishii, Koike, Kitagawa, Kimura, Imoto, Miyano, Ogawa, Kanai, Fukunaga and Okada2022); OR 0.78 for O vs. A, OR 0.79 for O vs. B, OR 0.65 for O vs. AB for individuals of European ancestry from USA and United Kingdom (Shelton et al., Reference Shelton, Shastri, Ye, Weldon, Filshtein-Sonmez, Coker, Symons, Esparza-Gordillo, Aslibekyan and Auton2021); OR 0.64 for O vs. A, OR 0.51 for O vs. B for Latin Americans and OR 0.43 for O vs. B for African Americans (Shelton et al., Reference Shelton, Shastri, Ye, Weldon, Filshtein-Sonmez, Coker, Symons, Esparza-Gordillo, Aslibekyan and Auton2021), sometimes inconsistent results from other studies (Leaf et al., Reference Leaf, Al-Samkari, Brenner, Gupta and Leaf2020)), the actual effect or impact of these associations at the biological level and for the disease process in COVID-19 is difficult to assess. The impact on infection rate may depend strongly on the underlying pathophysiological mechanism, the ABO blood group distribution in the population of interest, socioeconomic interventions in different countries, and the proportion of the population already infected at a given time, among other factors. Interestingly, contrary to initial studies (Zietz et al., Reference Zietz, Zucker and Tatonetti2020), no association with the RHD locus was found (Shelton et al., Reference Shelton, Shastri, Ye, Weldon, Filshtein-Sonmez, Coker, Symons, Esparza-Gordillo, Aslibekyan and Auton2021), suggesting that the rhesus factor on its own (and independent of the ABO blood group) is not a genetic risk factor. No significant difference was also found between the rhesus-positive and rhesus-negative forms of each ABO blood group.

Figure 1. Summary of results from meta-analysis association studies at the 9q34.2 locus (ABO) conducted by the COVID-19 Host Genetics Initiative (HGI). Meta-analyses of association data show an association of the 9q34.2 locus (ABO) with (i) critical severity of illness, (ii) hospitalization and (iii) infection, as described in COVID-19 Host Genetics Initiative (2022). Upper Manhattan plot: association results for 8,779 critically ill COVID-19 patients versus 1,001,875 population controls. Middle Manhattan plot: association results for 24,274 hospitalized COVID-19 patients versus 2,061,529 population controls. Lower Manhattan plot: association results for 112,612 SARS-CoV-2 infected individuals versus 2,474,079 population controls. X-axis: chromosome positions and gene annotations on human genome build hg38. Y-axis: meta-analysis association p-values (−log10p) of genetic markers. Plots were generated with the COVID-19 Host Genetics Initiative Browser (https://app.covid19hg.org; release 6).
The inclusion of controls with unknown status in most GWAS on disease severity may have led to associations with disease severity also being related to infection, as hospitalized cases are susceptible to infection, but an untested control group may or may not be susceptible, so it would be helpful to test against asymptomatic cases as well. Investigation of possible associations with severity is also possible with a comparison of blood group frequencies between patient subgroups with different clinical characteristics; results of studies with patient subgroups (not exhaustive) are listed in Table 3. These studies have further shown that blood group O is associated with lower disease severity, which is consistent with previously reported (SARS-CoV-2 independent) effects of ABO blood groups on thrombosis and vascular function (Vasan et al., Reference Vasan, Rostgaard, Majeed, Ullum, Titlestad, Pedersen, Erikstrup, Nielsen, Melbye, Nyren, Hjalgrim and Edgren2016). Thus, numerous studies indicate that the ABO locus and ABO blood groups are susceptibility factors for SARS-CoV-2 infection and COVID-19 severity.
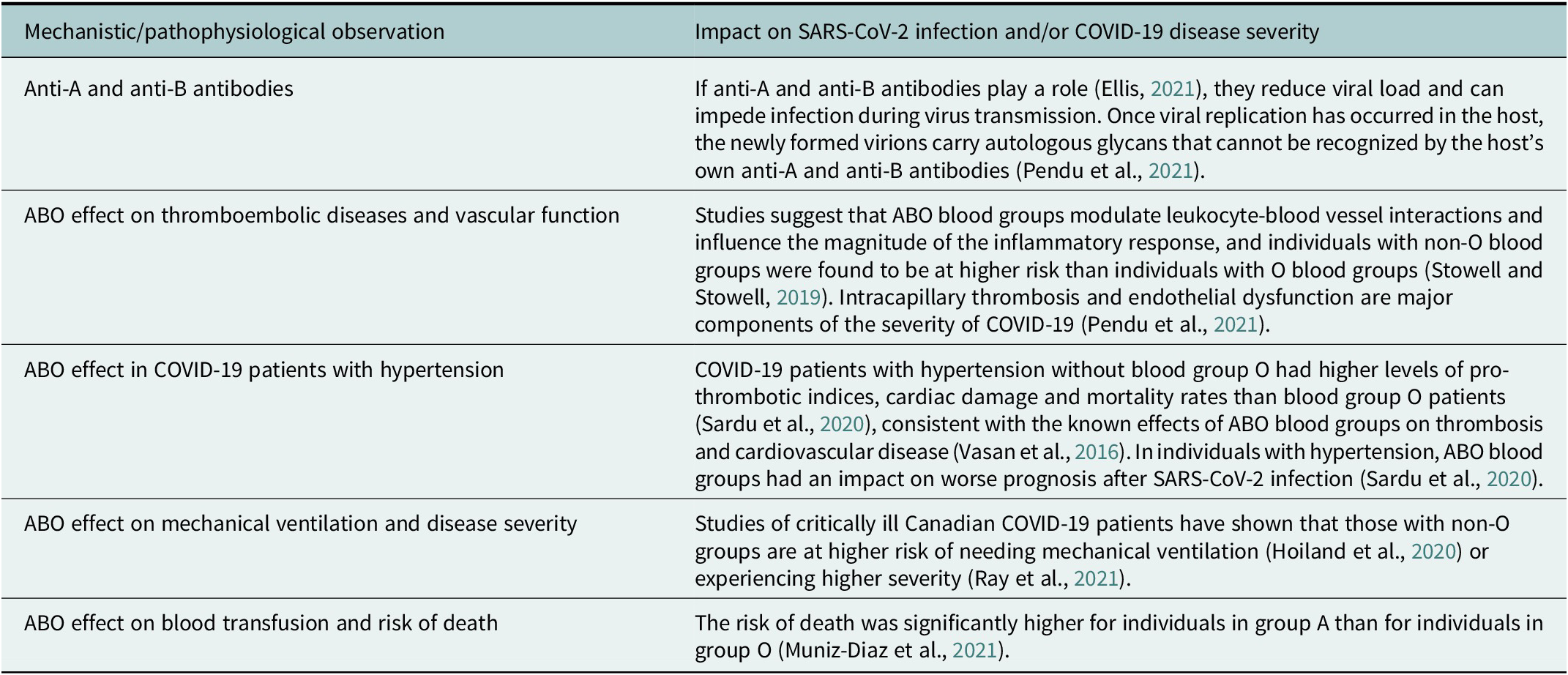
Table 3. Studies of mechanistic and pathophysiological hypotheses of ABO blood group effects as well as clinical findings from COVID-19 patient subgroup studies (not exhaustive) suggest an association between ABO blood groups and SARS-CoV-2 infection and COVID-19 disease severity
SARS-CoV-2 transmission models in the context of ABO blood group effects
Two potential pathophysiological mechanisms (Figure 2) have mainly been proposed to explain the reported association between ABO blood groups and the risk of SARS-CoV-2 infection: The ABO compatibility-dependence (or ABO-interference) hypothesis (neutralization by natural anti-ABO antibodies), as previously described for SARS-CoV-1 (Breiman et al., Reference Breiman, Ruven-Clouet and Le Pendu2020), and the ABO-dependent intrinsic hypothesis (direct attachment of the virus spike protein to blood group A glycans), as previously described for noroviruses and rotaviruses (Le Pendu and Ruvoen-Clouet, Reference Le Pendu and Ruvoen-Clouet2020).

Figure 2. Two predominant hypotheses of possible mechanisms involving ABO blood group-related antigens: (A) The ABO-compatibility-dependence model (or ABO-interference) and (B) the ABO-dependent intrinsic model. The ABO-compatibility dependence model was recently modeled by Ellis (Reference Ellis2021) under different assumptions and compared with observational healthcare data (Zietz et al., Reference Zietz, Zucker and Tatonetti2020; Zhao et al., Reference Zhao, Yang, Huang, Li, Gu, Lu, Zhang, Liu, Liu, Liu, He, Sun, Wei, Yang, Wang, Zhang, Zhou, Xing and Wang2021) and GWAS data from the Severe COVID-19 GWAS Group (Ellinghaus et al., Reference Ellinghaus, Degenhardt, Bujanda, Buti, Albillos, Invernizzi, Fernandez, Prati, Baselli, Asselta, Grimsrud, Milani, Aziz, Kassens, May, Wendorff, Wienbrandt, Uellendahl-Werth, Zheng, Yi, de Pablo, Chercoles, Palom, Garcia-Fernandez, Rodriguez-Frias, Zanella, Bandera, Protti, Aghemo, Lleo, Biondi, Caballero-Garralda, Gori, Tanck, Carreras Nolla, Latiano, Fracanzani, Peschuck, Julia, Pesenti, Voza, Jimenez, Mateos, Jimenez, Quereda, Paccapelo, Gassner, Angelini, Cea, Solier, Pestana, Muniz-Diaz, Sandoval, Paraboschi, Navas, Garcia Sanchez, Ceriotti, Martinelli-Boneschi, Peyvandi, Blasi, Tellez, Blanco-Grau, Hemmrich-Stanisak, Grasselli, Costantino, Cardamone, Foti, Aneli, Kurihara, ElAbd, My, Galvan-Femenia, Martin, Erdmann, Ferrusquia-Acosta, Garcia-Etxebarria, Izquierdo-Sanchez, Bettini, Sumoy, Terranova, Moreira, Santoro, Scudeller, Mesonero, Roade, Ruhlemann, Schaefer, Carrabba, Riveiro-Barciela, Basso, Valsecchi, Hernandez-Tejero, Acosta-Herrera, D’Angio, Baldini, Cazzaniga, Schulzky, Cecconi, Wittig, Ciccarelli, Rodriguez-Gandia, Bocciolone, Miozzo, Montano, Braun, Sacchi, Martinez, Ozer, Palmieri, Faverio, Preatoni, Bonfanti, Omodei, Tentorio, Castro, Rodrigues, Ortiz, de Cid, Ferrer, Gualtierotti, Nieto, Goerg, Badalamenti, Marsal, Matullo, Pelusi, Juzenas, Aliberti, Monzani, Moreno, Wesse, Lenz, Pumarola, Rimoldi, Bosari, Albrecht, Peter, Romero-Gomez, D’Amato, Duga, Banales, Hov, Folseraas, Valenti, Franke and Karlsen2020). Both models have been further evaluated by Boukhari et al. (Reference Boukhari, Breiman, Jazat, Ruvoen-Clouet, Martinez, Damais-Cepitelli, Le Niger, Devie-Hubert, Penasse, Mauriere, Sebille, Durrbach and Le Pendu2021) in a French study population of 666 individuals (333 index persons and their spouses) of known ABO blood type with a high risk of SARS-CoV-2 transmission (hospital employees) as well as receptor-binding domain (RBD) protein binding experiments in cell lines and saliva samples from individuals of known ABO and secretor phenotypes. For the ABO-compatibility-dependence model, ρ represents the relative probability of virus transmission between an infected index person and an ABO-incompatible contact (impeded transmission; pairs denoted with “I”) and was estimated to be 40% on average (between 20 and 55% depending on ABO blood group frequencies and relative risk ratios in different countries) by Ellis (Reference Ellis2021). Boukhari et al. estimated a decrease of 41% in ABO-incompatible pairs. The ABO-dependent intrinsic hypothesis remains controversial because of conflicting study results (Boukhari et al., Reference Boukhari, Breiman, Jazat, Ruvoen-Clouet, Martinez, Damais-Cepitelli, Le Niger, Devie-Hubert, Penasse, Mauriere, Sebille, Durrbach and Le Pendu2021; Wu et al., Reference Wu, Arthur, Wang, Verkerke, Josephson, Kalman, Roback, Cummings and Stowell2021). n.a., not available. Figure based on Boukhari et al. (Reference Boukhari, Breiman, Jazat, Ruvoen-Clouet, Martinez, Damais-Cepitelli, Le Niger, Devie-Hubert, Penasse, Mauriere, Sebille, Durrbach and Le Pendu2021) and extended.
If SARS-CoV-2 viruses replicate in respiratory tract cells that express ABO antigens (depending on the host’s ABO blood group and secretor status) then the A, B or H epitopes may also be present on the viral envelope glycoproteins due to the host cell glycosyltransferases, as shown in in vitro studies (Deleers et al., Reference Deleers, Breiman, Daubie, Maggetto, Barreau, Besse, Clemenceau, Ruvoen-Clouet, Fils, Maillart, Doyen, Mahadeb, Jani, Van der Linden, Cannie, Hayef, Corazza, Le Pendu and El Kenz2021). Thus, analogous to the rules of blood transfusion, in ABO incompatible situations (denoted with “I” in Figure 2A) of an index person (the one who transmits the virus) and a contact person (the one who receives the virus), we can speculate that the transmitted SARS-CoV-2 virus particles are neutralized by anti-A and anti-B antibodies, in which case individuals of blood group 0 would be at an advantage because they have both anti-A and anti-B antibodies. This hypothesis was first supported by previous in vitro observations for SARS-CoV-1 that anti-A antibodies can specifically block the interaction between the SARS-CoV-1 spike (S)-glycoprotein and its target, the ACE2 receptor (Guillon et al., Reference Guillon, Clement, Sebille, Rivain, Chou, Ruvoen-Clouet and Le Pendu2008). Therefore, early after the outbreak of the COVID-19 pandemic, Breiman et al. hypothesized that in the presence of sufficient anti-A and/or anti-B antibody titers, individuals with blood groups O, A and B might also have some protection against transmission of SARS-CoV-2 by infected ABO-incompatible persons (Breiman et al., Reference Breiman, Ruven-Clouet and Le Pendu2020). This hypothesis has now been explored by Ellis (Reference Ellis2021) using more refined modeling techniques and COVID-19 GWAS (Ellinghaus et al., Reference Ellinghaus, Degenhardt, Bujanda, Buti, Albillos, Invernizzi, Fernandez, Prati, Baselli, Asselta, Grimsrud, Milani, Aziz, Kassens, May, Wendorff, Wienbrandt, Uellendahl-Werth, Zheng, Yi, de Pablo, Chercoles, Palom, Garcia-Fernandez, Rodriguez-Frias, Zanella, Bandera, Protti, Aghemo, Lleo, Biondi, Caballero-Garralda, Gori, Tanck, Carreras Nolla, Latiano, Fracanzani, Peschuck, Julia, Pesenti, Voza, Jimenez, Mateos, Jimenez, Quereda, Paccapelo, Gassner, Angelini, Cea, Solier, Pestana, Muniz-Diaz, Sandoval, Paraboschi, Navas, Garcia Sanchez, Ceriotti, Martinelli-Boneschi, Peyvandi, Blasi, Tellez, Blanco-Grau, Hemmrich-Stanisak, Grasselli, Costantino, Cardamone, Foti, Aneli, Kurihara, ElAbd, My, Galvan-Femenia, Martin, Erdmann, Ferrusquia-Acosta, Garcia-Etxebarria, Izquierdo-Sanchez, Bettini, Sumoy, Terranova, Moreira, Santoro, Scudeller, Mesonero, Roade, Ruhlemann, Schaefer, Carrabba, Riveiro-Barciela, Basso, Valsecchi, Hernandez-Tejero, Acosta-Herrera, D’Angio, Baldini, Cazzaniga, Schulzky, Cecconi, Wittig, Ciccarelli, Rodriguez-Gandia, Bocciolone, Miozzo, Montano, Braun, Sacchi, Martinez, Ozer, Palmieri, Faverio, Preatoni, Bonfanti, Omodei, Tentorio, Castro, Rodrigues, Ortiz, de Cid, Ferrer, Gualtierotti, Nieto, Goerg, Badalamenti, Marsal, Matullo, Pelusi, Juzenas, Aliberti, Monzani, Moreno, Wesse, Lenz, Pumarola, Rimoldi, Bosari, Albrecht, Peter, Romero-Gomez, D’Amato, Duga, Banales, Hov, Folseraas, Valenti, Franke and Karlsen2020) and prevalence data (Zietz et al., Reference Zietz, Zucker and Tatonetti2020; Zhao et al., Reference Zhao, Yang, Huang, Li, Gu, Lu, Zhang, Liu, Liu, Liu, He, Sun, Wei, Yang, Wang, Zhang, Zhou, Xing and Wang2021) from regions in the early phase of the SARS-CoV-2 epidemic. If the model holds, ABO incompatibility reduces viral transmissibility by 60% (Ellis, Reference Ellis2021), but the relative risk for each blood group is nearly the same once the majority of a given population is infected (see French Navy aircraft study (Boudin et al., Reference Boudin, Janvier, Bylicki and Dutasta2020) below).
In contrast, according to the ABO-dependent intrinsic hypothesis (Figure 2B), individuals with blood groups A, B and AB are inherently more susceptible to SARS-CoV-2 infection than individuals with blood group O, regardless of the blood group of the transmitting index person. Only the blood group of the contact person plays a role here. The difference in susceptibility is attributed to the possibility of direct binding of the receptor-binding domain (RBD) of the SARS-CoV-2 spike protein to blood group glycans (such as the A antigen) (Wu et al., Reference Wu, Arthur, Wang, Verkerke, Josephson, Kalman, Roback, Cummings and Stowell2021), which could facilitate the infection process and make individuals with non-O blood groups more vulnerable. Similar results were obtained from human noroviruses and rotaviruses studies, where the absence or low expression of the recognized glycan motifs due to combined ABO, FUT2 and FUT3 gene polymorphisms was associated with resistance to the diarrheal disease (Le Pendu and Ruvoen-Clouet, Reference Le Pendu and Ruvoen-Clouet2020).
In favor of the ABO-compatibility-dependence model
The effects of the two hypotheses could potentially overlap, but with different consequences for different constellations of ABO blood group frequencies in the population. In an attempt to test the two hypotheses based on further data, Boukhari et al. (Reference Boukhari, Breiman, Jazat, Ruvoen-Clouet, Martinez, Damais-Cepitelli, Le Niger, Devie-Hubert, Penasse, Mauriere, Sebille, Durrbach and Le Pendu2021) calculated symptomatic secondary attack rates (SAR) in 333 French couples (333 index persons (hospital employees) and their spouses who constantly slept in the same bedroom; only symptomatic infections within 7 days with PCR confirmation were counted) with known blood groups and assessed the effect of ABO compatibility/incompatibility. This study was convincing because of its very good study design; some previous ABO blood group studies on other infections may have lacked such a design, which could have led to inconclusive results. In addition, they experimentally reinvestigated the reported potential binding of the SARS-CoV-2 RBD to blood group A epitopes (Wu et al., Reference Wu, Arthur, Wang, Verkerke, Josephson, Kalman, Roback, Cummings and Stowell2021).
ABO incompatibility was significantly associated with a lower risk of symptomatic COVID-19 transmission (P = 0.0004; OR 0.43, 95% CI 0.27–0.69); the SAR was 47.2% in ABO-compatible couples, but only 27.9% in ABO-incompatible couples (decrease of 41%). After further classification of ABO-compatible and ABO-incompatible couples according to the ABO blood group of the contact partner (spouse), SAR were always higher in ABO-compatible couples than in ABO-incompatible couples, regardless of the blood group of the contact partner (with exception of blood group AB, which cannot be incompatible, see Figure 2A). This means that the risk of transmission to spouses of blood group A was not higher than that to spouses of other blood groups with ABO compatibility, suggesting that all ABO blood groups are intrinsically equally susceptible to COVID-19 (contradictory result to the hypothesis in Figure 2B and previous studies (Wu et al., Reference Wu, Arthur, Wang, Verkerke, Josephson, Kalman, Roback, Cummings and Stowell2021)). Interestingly, as in the GWAS studies, the COVID-19 negative subgroup had a much higher frequency of blood group O and a much lower frequency of blood group A compared to the COVID-19 positive subgroup. In their binding experiments, the authors were further unable to demonstrate, among other things, either binding of the recombinant chimeric RBD-Fc protein to the blood group A 1 type chain structures (expressed on epithelial cells) or binding above background level (in dependence of the ABO blood group of the donor) of the RBD to salivary mucins from saliva samples (salivary mucins contain histo-blood group antigens similar to those in epithelia; saliva samples with secretor phenotype were selected to ensure expression of A, B or H(O) histo-blood group antigens).
The impact of the ABO-compatibility-dependence model on the population and individual level
Different blood group antigens can generally increase or decrease host susceptibility to infections (Cooling, Reference Cooling2015). The two most common ABO blood groups in Western Europe are blood group A and blood group O. On the Eurasian continent, there is a gradient of blood group B, whose frequency increases from west to east (where a more balanced distribution between all ABO blood groups dominates and where also, interestingly, an ancient viral selection pressure on host coronavirus interacting genes began more than 20,000 years ago (Souilmi et al., Reference Souilmi, Lauterbur, Tobler, Huber, Johar, Moradi, Johnston, Krogan, Alexandrov and Enard2021)). In Africa, blood group O is the most common (Cooling, Reference Cooling2015), and the Americas show the high proportion of blood type O (see Cabezas-Cruz et al., Reference Cabezas-Cruz, Mateos-Hernandez, Alberdi, Villar, Riveau, Hermann, Schacht, Khalife, Correia-Neves, Gortazar and de la Fuente2017, Supplementary Table 1, for an excellent overview of published geographic ABO blood group distributions worldwide or https://biobankengine.shinyapps.io/hla-map/, tab “ABO Global Map”). On the population level, according to the ABO-compatibility-dependence hypothesis, transmission rates then would have to be higher in populations with a high frequency of blood group O than in populations in which this blood type is less common because the frequency of compatible encounters is then higher. Indeed, Pendu et al. (Reference Pendu, Breiman, Rocher, Dion and Ruvoen-Clouet2021) demonstrated (ruling out to a certain extent confounder effects due to socioeconomic and demographic inequalities and differences in protective measures among various countries) that the estimated protective effect of blood type O over A/B/AB was significantly higher in populations with blood group percentages below 40% than in populations with blood group percentages above 40%, and that countries where blood group O is most prevalent also had the highest SARS-CoV-2 infection rates. On an individual level, however, it is very likely that more or less no one can escape SARS-CoV-2 infection if a large part of the population is already infected. This can be illustrated by the isolated outbreak of SARS-CoV-2 in 2020, well-known from the media, when 75.8% of 1,769 crew members of the aircraft carrier Charles de Gaulle of the French Navy became infected, but no statistical association between ABO blood groups and SARS-CoV-2 incidence was found (Boudin et al., Reference Boudin, Janvier, Bylicki and Dutasta2020).
Conclusion and future perspectives
Twenty-five susceptibility loci for SARS-CoV-2 infection and/or COVID-19 have been identified through GWAS, and the most frequently replicated genetic finding is the ABO gene on chromosome 9q34.2, which encodes glycosyltransferases important for A and B antigens on epithelial cells, such as those of the upper respiratory tract and the digestive tract. The analysis of secondary attack rates (SAR) in ABO blood group incompatible pairs of individuals (denoted by pairs “I” in Figure 2A) as well as SARS-CoV-2 RBD experiments revealed no evidence for the ABO-dependent intrinsic susceptibility hypothesis (Figure 2B, i.e., higher susceptibility for blood group A due to potential binding of the SARS-CoV-2 RBD to the A antigen). Instead, the ABO-compatibility-dependence model is currently favored, where transmission in ABO incompatibility situations were associated with a much lower SAR than transmission in ABO compatibility situations (denoted by pairs “U” in Figure 2A). The results suggest that natural anti-AB antibodies can reduce the risk of SARS-CoV-2 transmission by up to 41% in blood group 0 individuals (Boukhari et al., Reference Boukhari, Breiman, Jazat, Ruvoen-Clouet, Martinez, Damais-Cepitelli, Le Niger, Devie-Hubert, Penasse, Mauriere, Sebille, Durrbach and Le Pendu2021), consistent with a value of 40% (Ellis, Reference Ellis2021) (20–55% depending on ABO blood group frequencies and relative risk ratios in different countries) estimated from COVID-19 GWAS and observational healthcare data (Ellinghaus et al., Reference Ellinghaus, Degenhardt, Bujanda, Buti, Albillos, Invernizzi, Fernandez, Prati, Baselli, Asselta, Grimsrud, Milani, Aziz, Kassens, May, Wendorff, Wienbrandt, Uellendahl-Werth, Zheng, Yi, de Pablo, Chercoles, Palom, Garcia-Fernandez, Rodriguez-Frias, Zanella, Bandera, Protti, Aghemo, Lleo, Biondi, Caballero-Garralda, Gori, Tanck, Carreras Nolla, Latiano, Fracanzani, Peschuck, Julia, Pesenti, Voza, Jimenez, Mateos, Jimenez, Quereda, Paccapelo, Gassner, Angelini, Cea, Solier, Pestana, Muniz-Diaz, Sandoval, Paraboschi, Navas, Garcia Sanchez, Ceriotti, Martinelli-Boneschi, Peyvandi, Blasi, Tellez, Blanco-Grau, Hemmrich-Stanisak, Grasselli, Costantino, Cardamone, Foti, Aneli, Kurihara, ElAbd, My, Galvan-Femenia, Martin, Erdmann, Ferrusquia-Acosta, Garcia-Etxebarria, Izquierdo-Sanchez, Bettini, Sumoy, Terranova, Moreira, Santoro, Scudeller, Mesonero, Roade, Ruhlemann, Schaefer, Carrabba, Riveiro-Barciela, Basso, Valsecchi, Hernandez-Tejero, Acosta-Herrera, D’Angio, Baldini, Cazzaniga, Schulzky, Cecconi, Wittig, Ciccarelli, Rodriguez-Gandia, Bocciolone, Miozzo, Montano, Braun, Sacchi, Martinez, Ozer, Palmieri, Faverio, Preatoni, Bonfanti, Omodei, Tentorio, Castro, Rodrigues, Ortiz, de Cid, Ferrer, Gualtierotti, Nieto, Goerg, Badalamenti, Marsal, Matullo, Pelusi, Juzenas, Aliberti, Monzani, Moreno, Wesse, Lenz, Pumarola, Rimoldi, Bosari, Albrecht, Peter, Romero-Gomez, D’Amato, Duga, Banales, Hov, Folseraas, Valenti, Franke and Karlsen2020; Zietz et al., Reference Zietz, Zucker and Tatonetti2020; Zhao et al., Reference Zhao, Yang, Huang, Li, Gu, Lu, Zhang, Liu, Liu, Liu, He, Sun, Wei, Yang, Wang, Zhang, Zhou, Xing and Wang2021). However, these estimates can only be valid if a large portion of the population is not yet infected with SARS-CoV-2. In previous studies, it has been reported that women with blood group O have higher natural anti-A antibody levels than men with blood type O and that antibody titers decrease with increasing age (de Franca et al., Reference de Franca, Poli, Ramos, Borsoi and Colella2011; McVey et al., Reference McVey, Baker, Parti, Berg, Gudino and Teschner2015). Therefore, future models could also account for possible sex- and age-related differences in SARS-CoV-2 transmission.
Major efforts by the worldwide GWAS community to unravel the genetic basis of COVID-19 have consistently identified genes with high biological plausibility, but no convincing polygenic risk score (PRS) for clinical testing for COVID-19 severity/SARS-CoV-2 infection has yet been published. This is a general problem with GWAS studies (Wald and Old, Reference Wald and Old2019), which is why PRS will be more useful for stratifying patients into subgroups. However, drug repurposing may offer a rapid approach to address the urgent need for therapeutics for COVID-19. Some of the genes identified in the COVID-19 GWAS studies, for example, IFNAR2 and ACE2, encode proteins against which drug candidates are currently being tested in clinical trials (Gaziano et al., Reference Gaziano, Giambartolomei, Pereira, Gaulton, Posner, Swanson, Ho, Iyengar, Kosik, Vujkovic, Gagnon, Bento, Barrio-Hernandez, Ronnblom, Hagberg, Lundtoft, Langenberg, Pietzner, Valentine, Gustincich, Tartaglia, Allara, Surendran, Burgess, Zhao, Peters, Prins, Angelantonio, Devineni, Shi, Lynch, DuVall, Garcon, Thomann, Zhou, Gorman, Huffman, O’Donnell, Tsao, Beckham, Pyarajan, Muralidhar, Huang, Ramoni, Beltrao, Danesh, Hung, Chang, Sun, Joseph, Leach, Edwards, Cho, Gaziano, Butterworth and Casas2021). In addition, GWAS studies of other COVID-19 symptoms such as loss of smell or taste (Shelton et al., Reference Shelton, Shastri, Fletez-Brant, Aslibekyan and Auton2022) could help elucidate the underlying biology, which could be further steps toward new precise treatments. It remains to be seen whether the genetic associations in COVID-19 can make a significant direct contribution to precision medicine. However, it should be remembered that genetic studies in patients are one of the best ways to explore new therapeutic targets for precision medicine: The overall added value is that GWAS conduct genetic research directly in humans, allowing us to study and uncover individual patient risk factors in a hypothesis-free manner. It has been estimated that selecting genetically based targets generally doubles the success rate in clinical development (Nelson et al., Reference Nelson, Tipney, Painter, Shen, Nicoletti, Shen, Floratos, Sham, Li, Wang, Cardon, Whittaker and Sanseau2015).
Not only various host factors such as sex, age, genetics and comorbidities, but also environmental factors such as the gut microbiome could have a major impact on SARS-CoV-2 infection and disease severity in COVID-19 (Chhibber-Goel et al., Reference Chhibber-Goel, Gopinathan and Sharma2021; Yeoh et al., Reference Yeoh, Zuo, Lui, Zhang, Liu, Li, Chung, Cheung, Tso, Fung, Chan, Ling, Joynt, Hui, Chow, Ng, Li, Ng, Yip, Wong, Chan, Wong, Chan and Ng2021; Wang et al., Reference Wang, Zhang, Wang, Dai, Qin, Zhou and Zhang2022). The composition of the microbiota varies between individuals and populations, and the composition of the gut microbiota is also genetically influenced (Kurilshikov et al., Reference Kurilshikov, Medina-Gomez, Bacigalupe, Radjabzadeh, Wang, Demirkan, Le Roy, Raygoza Garay, Finnicum, Liu, Zhernakova, Bonder, Hansen, Frost, Ruhlemann, Turpin, Moon, Kim, Lull, Barkan, Shah, Fornage, Szopinska-Tokov, Wallen, Borisevich, Agreus, Andreasson, Bang, Bedrani, Bell, Bisgaard, Boehnke, Boomsma, Burk, Claringbould, Croitoru, Davies, van Duijn, Duijts, Falony, Fu, van der Graaf, Hansen, Homuth, Hughes, Ijzerman, Jackson, Jaddoe, Joossens, Jorgensen, Keszthelyi, Knight, Laakso, Laudes, Launer, Lieb, Lusis, Masclee, Moll, Mujagic, Qibin, Rothschild, Shin, Sorensen, Steves, Thorsen, Timpson, Tito, Vieira-Silva, Volker, Volzke, Vosa, Wade, Walter, Watanabe, Weiss, Weiss, Weissbrod, Westra, Willemsen, Payami, Jonkers, Vasquez, de Geus, Meyer, Stokholm, Segal, Org, Wijmenga, Kim, Kaplan, Spector, Uitterlinden, Rivadeneira, Franke, Lerch, Franke, Sanna, D’Amato, Pedersen, Paterson, Kraaij, Raes and Zhernakova2021). ABO blood groups and Secretor/FUT2 status have already been associated with gut microbiome characteristics (Ruhlemann et al., Reference Ruhlemann, Hermes, Bang, Doms, Moitinho-Silva, Thingholm, Frost, Degenhardt, Wittig, Kassens, Weiss, Peters, Neuhaus, Volker, Volzke, Homuth, Weiss, Grallert, Laudes, Lieb, Haller, Lerch, Baines and Franke2021). Several studies have shown that SARS-CoV-2 infection has negative effects on the respiratory, intestinal and oral microbiota (Gang et al., Reference Gang, Wang, Xue and Zhang2022), manifested mainly in a decrease in microbial diversity and beneficial symbiotic bacteria and an increase in opportunistic pathogens (Zuo et al., Reference Zuo, Zhang, Lui, Yeoh, Li, Zhan, Wan, Chung, Cheung, Chen, Lai, Chen, Tso, Fung, Chan, Ling, Joynt, Hui, Chan, Chan and Ng2020; Gao et al., Reference Gao, Wang, Luo, Sun, Wang, Ding, Ren, Gang, Rao, Liu, Wang, Gao, Li, Zou, Liu, Yuan, Sun, Cui and Ren2021; Ren et al., Reference Ren, Wang, Cui, Lu, Wang, Luo, Chen, Ren, Sun, Liu, Liu, Liu, Li, Wang, Rao, Yuan, Zhang, Sun, Chen, Li, Hu, Wu, Yu, Kan and Li2021) or by priming of host inflammatory responses by the gut microbiome and regulation of endocrine signaling (Sarkar et al., Reference Sarkar, Harty, Moeller, Klein, Erdman, Friston and Carmody2021); however, studies on the effects of ABO blood groups on the human microbiome in COVID-19 patients are lacking. The extent to which the observed ABO blood group effects also apply to novel SARS-CoV-2 viral variants (as of 2022) is unknown. Whether the ABO-compatibility-dependence model also shows the same effects in people who have undergone reinfection with SARS-CoV-2 has not yet been investigated. According to the latest study results (infections until 2021; without Omicron variants), the risk for long-COVID is still present after previous vaccination (breakthrough SARS-CoV-2 infection) or could even increase after reinfections (Al-Aly et al., Reference Al-Aly, Bowe and Xie2022a). Whether the risk of COVID-19 increases with reinfections with SARS-CoV-2 is unclear, and recent study results sometimes contradict each other (Abu-Raddad et al., Reference Abu-Raddad, Chemaitelly and Bertollini2021; Chemaitelly et al., Reference Chemaitelly, Bertollini and Abu-Raddad2021; Al-Aly et al., Reference Al-Aly, Bowe and Xie2022b). Therefore, future efforts to study blood group effects in the context of the human microbiome and other demographic and genetic variables, and in the context of multiple infections and newer viral variants, may provide more detailed insights into susceptibility to SARS-CoV-2 infection and the severity of COVID-19 in the context of ABO blood group effects.
Open peer review
To view the open peer review materials for this article, please visit http://doi.org/10.1017/pcm.2022.12.
Acknowledgments
Figure 2 was created with BioRender.com.
Financial support
This study was supported by the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation) Cluster of Excellence 2167 “Precision Medicine in Chronic Inflammation (PMI)” (EXC 2167-390884018).
Competing interest
The author declares no competing interests.










Comments
Dear Editors of Cambridge Prisms: Precision Medicine,
Thank you very much for inviting me to write a review article on "COVID-19 host genetics and ABO blood group susceptibility". Please find attached the review article you requested. I look forward to your response and I am happy to answer any further questions you may have.
Yours sincerely,
David Ellinghaus