Introduction
Species in Mermithidae Braun, 1883, one of two families in the nematode order Mermithida, have a pharynx posteriorly with a multicellular organ (stichosome) comprising longitudinal rows of gland-like cells called stichocytes (Maggenti, Reference Maggenti1981; Tchesunov & Spiridorov, Reference Tchesunov and Hope1993). The mermithid life cycle contains a parasitic juvenile stage and a free-living adult stage (see Poinar & Mullens, Reference Poinar and Mullens1987). More than 500 mermithid species have been reported (Nemys, 2022), most of which use insects as their hosts, but with a few using non-insect hosts such as slugs (Mollusca: Gastropoda), nematodes, spiders (Chelicerata: Araneae) or crustaceans (e.g. Vandergast & Roderick, Reference Vandergast and Roderick2003; Ross et al., Reference Ross, Ivanova, Spiridonov, Waeyenberge, Moens, Nicol and Wilson2010; Holovachov & Boström, Reference Holovachov and Boström2013; Warren et al., Reference Warren, Dutton, Whelan, Yanong and Bullard2019). Nearly all mermithids have been reported from terrestrial or freshwater environments; only two species are known from the marine environment: Thalassomermis megamphis Tchesunov & Hope, 1997 and a species in Agamomermis (Tchesunov & Hope, Reference Tchesunov and Rozenberg1997; Dusto, Reference Dusto2020).
There is little information on the origin of marine mermithids. Like Mermithidae, the mermithidan family Tetradonematidae Cobb, 1919 likewise comprises mainly terrestrial/freshwater members. Recent phylogenetic studies have suggested a non-marine origin for Mermithida (e.g. Westerman et al., Reference Westerman, Neves, Ahmed and Holovachov2021), but no molecular data from marine mermithids have been included there.
Here, we report a mermithid species parasitic in a marine crustacean in the order Tanaidacea, which comprises about 1500 species (Anderson, Reference Anderson2020) and lacks any previous mermithid record. We provide an abbreviated description of the morphology of the nematode species, present nucleotide sequences for part of its nuclear 18S ribosomal RNA (18S) gene and infer the species’ phylogenetic position in the family Mermithidae based on 18S data.
Materials and methods
Tanaidaceans were collected by the R/V Yayoi (The University of Tokyo) with a biological dredge at 52 m depth in Otsuchi Bay (39°21.081′N, 141°58.145′E), Iwate, Japan, north-western Pacific on 28 April 2009, and preserved in 99% ethanol. Ethanol-preserved individuals were examined for parasite infection by ventral observation under an Olympus SZX9 stereomicroscope. The identification of hosts was based on Sieg (Reference Sieg1980), Bamber (Reference Bamber, Wells, Walker and Kendrick2005), Bamber & Boxshall (Reference Bamber and Boxshall2006), Kakui et al. (Reference Kakui, Katoh, Hiruta, Kobayashi and Kajihara2011), Larsen et al. (Reference Larsen, Guţu, Sieg, von Vaupel Klein, Charmantier-Daures and Schram2015) and Chim & Tong (Reference Chim and Tong2019).
To extract parasites, seven infected tanaidaceans were dissected with chemically sharpened needles under an SZX9 stereomicroscope; three host individuals bearing parasites were kept intact for future non-destructive observation. The anterior and/or posterior regions of three extracted parasites were transferred into a 1:3:6 mixture of glycerine, absolute ethanol and deionized water, and placed in a thermostatic chamber at 40°C for 4 h, after which they were mounted on glass slides in glycerine and observed with an Olympus BX51 microscope. Illustrations of parasites were prepared with Inkscape 1.0 (https://inkscape.org) from digital photomicrograph images captured with a PCM500 (AS ONE, Osaka, Japan) digital camera and edited with GIMP version 2.10 (https://www.gimp.org). The cephalothorax width (CW) of the tanaidaceans, as an indication of specimen size, was measured at the widest portion of the cephalothorax. The body length of one nematode individual for which we successfully retrieved all body fragments (see below) was measured from the anterior to posterior tips of the body. Ten infected tanaidaceans and extracted nematodes have been deposited in the Invertebrate Collection of the Hokkaido University Museum (ICHUM), Sapporo, under catalogue numbers ICHUM8290–ICHUM8299. The other tanaidaceans remain in the first author's private collection.
An attempt was made to extract total DNA from part of the body of each of the five nematodes obtained from five different host individuals using a NucleoSpin Tissue XS Kit (Macherey-Nagel, Düren, Germany), but only three of the five extracts allowed successful polymerase chain reaction (PCR) amplification. To obtain a nearly complete 18S sequence, two partly overlapping fragments were determined and then concatenated into a single fragment (cf. Holterman et al., Reference Holterman, van der Wurff, van den Elsen, van Megen, Bongers, Holovachov, Bakker and Helder2006). PCR primers for the first and second fragments were 1096F/1912R and 1813F/2646R (Holterman et al., Reference Holterman, van der Wurff, van den Elsen, van Megen, Bongers, Holovachov, Bakker and Helder2006), respectively; in addition, two primers, 18S-b3R (ATGTGGTAGCCGTTTCTCAGG, newly designed in this study) and 18S-b8F (Kakui et al., Reference Kakui, Katoh, Hiruta, Kobayashi and Kajihara2011), were used in cycle sequencing. PCR amplification conditions with KOD FX Neo polymerase (Toyobo, Osaka, Japan) were 94°C for 2 min; 45 cycles of 98°C for 10 s, 52°C (first fragment) or 55°C (second fragment) for 30 s and 68°C for 30 s; and 68°C for 2 min. All nucleotide sequences were determined by direct sequencing with a BigDye Terminator Kit version 3.1 (Thermo Fisher Scientific, Waltham, USA) with a 3730 DNA Analyzer (Thermo Fisher Scientific). Fragments were concatenated using MEGA7 (Kumar et al., Reference Kumar, Stecher and Tamura2016). The three 18S sequences we determined (which proved to be identical) were deposited in the International Nucleotide Sequence Database (INSD, 2022) through the DNA Data Bank of Japan, under accession numbers LC687375–LC687377.
The 18S dataset for a phylogenetic analysis included the unique sequence we determined and 58 sequences from 56 mermithidan species and two outgroup taxa (two species in Mononchida) taken from the INSD (table 1). The dataset was pre-aligned using MAFFT version 7 (Katoh & Standley, Reference Katoh and Standley2013) with the ‘Auto’ strategy, trimmed in MEGA7 to the shortest length among the sequences and then realigned by eye according to the secondary structure of the honeybee 18S sequence (Gillespie et al., Reference Gillespie, Johnston, Cannone and Gutell2006) and that of our used sequences predicted with the RNAfold WebServer (Gruber et al., Reference Gruber, Lorenz, Bernhart, Neuböck and Hofacker2008; Lorenz et al., Reference Lorenz, Bernhart, Zu Siederdissen, Tafer, Flamm, Stadler and Hofacker2011). The 687-bp aligned dataset used for phylogenetic reconstruction is presented in supplementary file S1. The optimal substitution model determined under the corrected Akaike information criterion option in ModelFinder (Kalyaanamoorthy et al., Reference Kalyaanamoorthy, Minh, Wong, von Haeseler and Jermiin2017) was TPM2 + F + I+G4. A maximum likelihood (ML) analysis was conducted in IQ-TREE version 2.1.2 (Minh et al., Reference Minh, Schmidt, Chernomor, Schrempf, Woodhams, von Haeseler and Lanfear2020); nodal support values were obtained from an ultrafast bootstrap analysis of 1000 pseudoreplicates under the ‘bnni’ option (Hoang et al., Reference Hoang, Chernomor, von Haeseler, Minh and Vinh2018). The ML tree was drawn with FigTree version 1.4.4 (Rambaut, Reference Rambaut2022).
Table 1. Information on the nematodes included in our phylogenetic analysis.
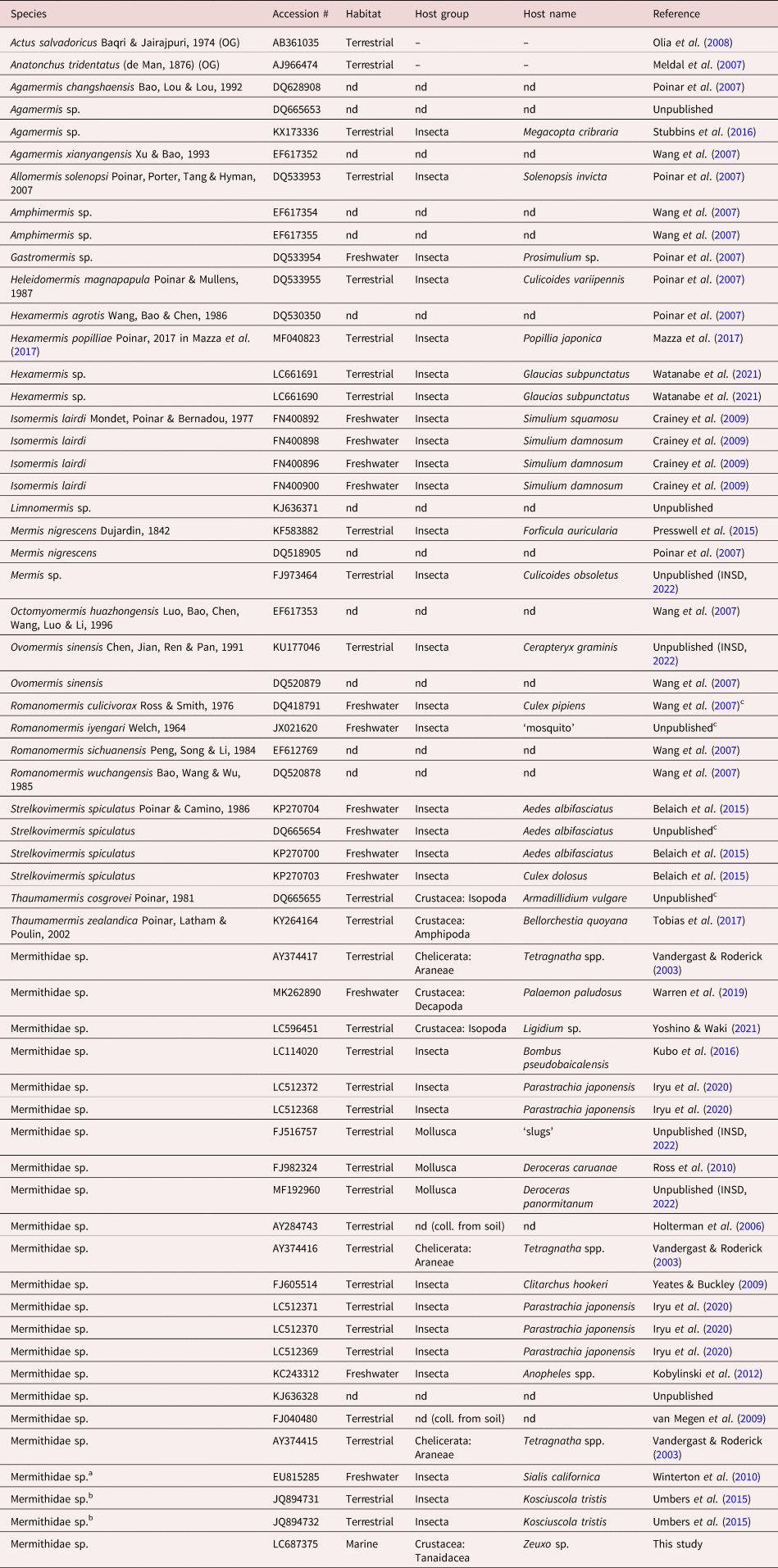
a This sequence was deposited as ‘Sialis californica’, but we regard it as coming from a mermithid nematode, based on the high identity score in BLAST searches with our sequence (query cover 90%; percent identity 91.30%).
b These sequences were deposited as ‘Nematoda sp.’, but we regard them as coming from mermithid nematodes, based on high identity scores in BLAST searches with our sequence (query cover 93–94%; percent identity 91.84–94.41%).
c Although no host data are presented in the INSD, one of the authors who deposited this sequence (Bradley Hyman) indicated this nematode species as extracted from this host species in other work(s) (see Hyman et al., Reference Hyman, Tang, Wu, Platzer and Pacheco2005; Pérez-Pacheco et al., Reference Pérez-Pacheco, Platzer, Woodward and Hyman2015).
OG, outgroup; nd, no data.
Results and discussion
We examined 599 tanaidaceans for parasite infection and found ten infected individuals (three males, CW 1.10–1.17 mm; seven females, CW 0.99–1.12 mm); some infected individuals may have been overlooked, as we did not perform dissections in our examination. We identified all tanaidaceans examined as the tanaidid Zeuxo sp. (fig. 1). Unfortunately, as the specimens had been preserved in absolute ethanol for more than ten years, hosts and parasites were strongly dehydrated and deformed, and the latter had become very fragile, with the result that all parasites were fragmented into two or more pieces after extraction (several pieces were damaged and lost). It was thus impossible for us to count the number of parasites per host. We did obtain two posterior portions from one host individual, indicating that a single host individual can bear multiple parasites.
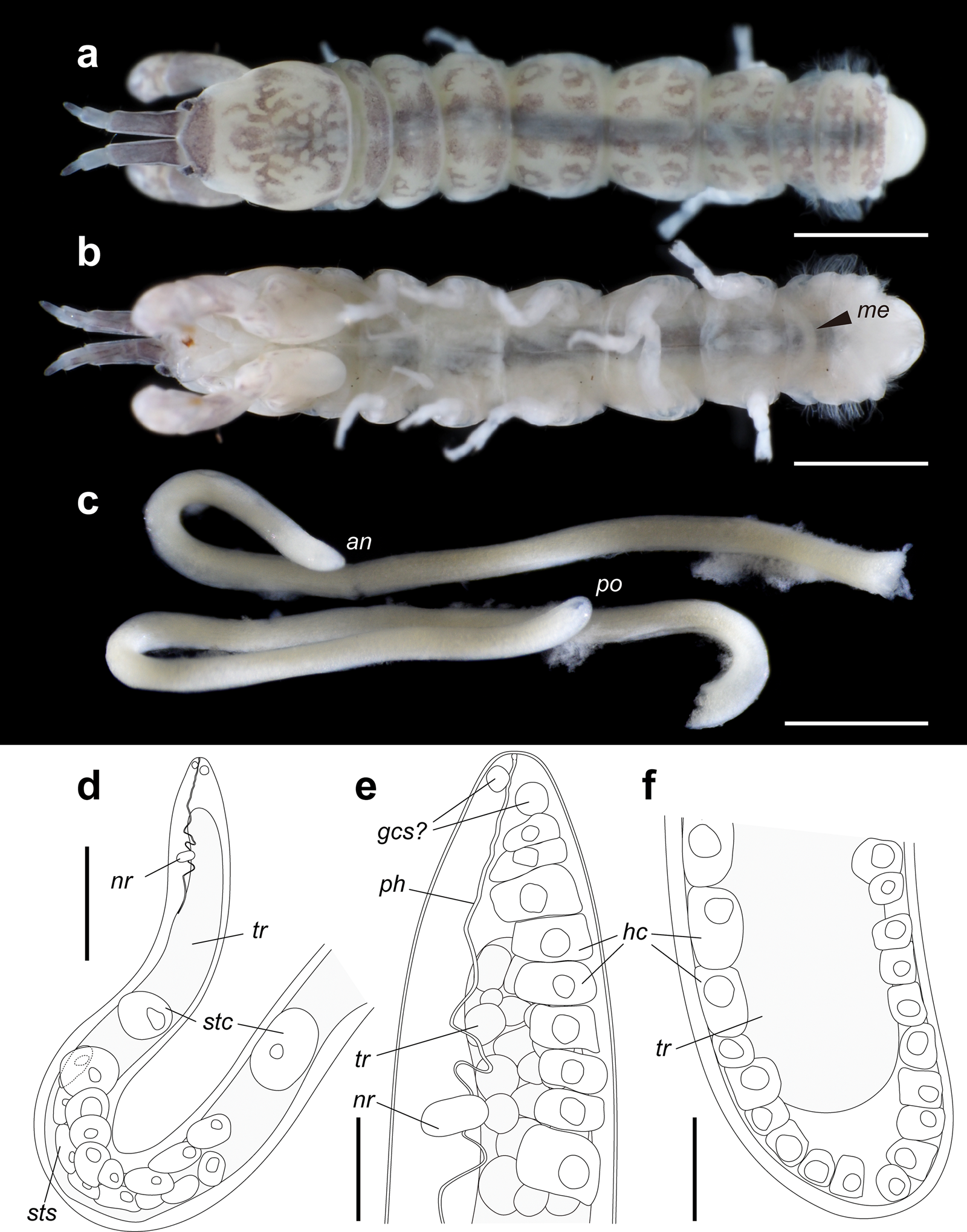
Fig. 1. Tanaidacean Zeuxo sp. and the infecting nematode parasite Mermithidae sp. (a, b) Infected Zeuxo sp., dorsal and ventral views, ethanol-preserved specimen (ICHUM8290). (c–f) Mermithidae sp., ethanol-preserved specimen (ICHUM8291); (c) photograph of anterior and posterior fragments; (d–f) anterior (d, e) and posterior (f) portions, line drawings. Scale bars: (a, b) 1 mm; (c) 0.5 mm; (d) 0.2 mm; (e, f) 0.05 mm. Abbreviations: an, anterior tip; gcs, gland cell of sensilla; hc, hypodermal cell; me, Mermithidae sp. inside the host; nr, nerve ring; ph, pharynx; po, posterior tip; stc, stichocyte; sts, stichosome; tr, trophosome.
The nematodes were found in the host body cavity. All body fragments were successfully retrieved in only one nematode individual (fig. 1c–f; ICHUM8291). An abbreviated description of this nematode is as follows. Body thread-like, 7.19 mm long. Maximum body diameter 135 μm. No transverse striation observed on cuticle. Dorsal, ventral and two lateral chords with single rows of well-developed hypodermal cells. Head rounded. Anterior sensilla and amphids not observed, but four (?) gland cells present in cephalic region. Buccal cavity cup-shaped, without stylet. Pharynx non-muscular, with cuticularized lumen, passing through nerve ring, posteriorly not connected to trophosome. Stichosome consisting of two rows of large stichocytes. Two single stichocytes present apart from stichosome. Trophosome filled with numerous granular cells and without lumen. Anus and rectum absent. Tail rounded.
The three 18S sequences we determined were identical and 1581 bp long. In the ML tree (fig. 2), they are nested in a well-supported clade (98% ultrafast bootstrap support (BS)) along with 13 sequences from other nematodes that infect terrestrial or freshwater hosts. In the tree, the nematode is the sister group (94% BS) to a fully supported clade comprising two nematodes that parasitized a spider (AY374417) and alderfly (EU815285). This result shows a shift in host utilization in Mermithidae from terrestrial or freshwater species to a marine species.
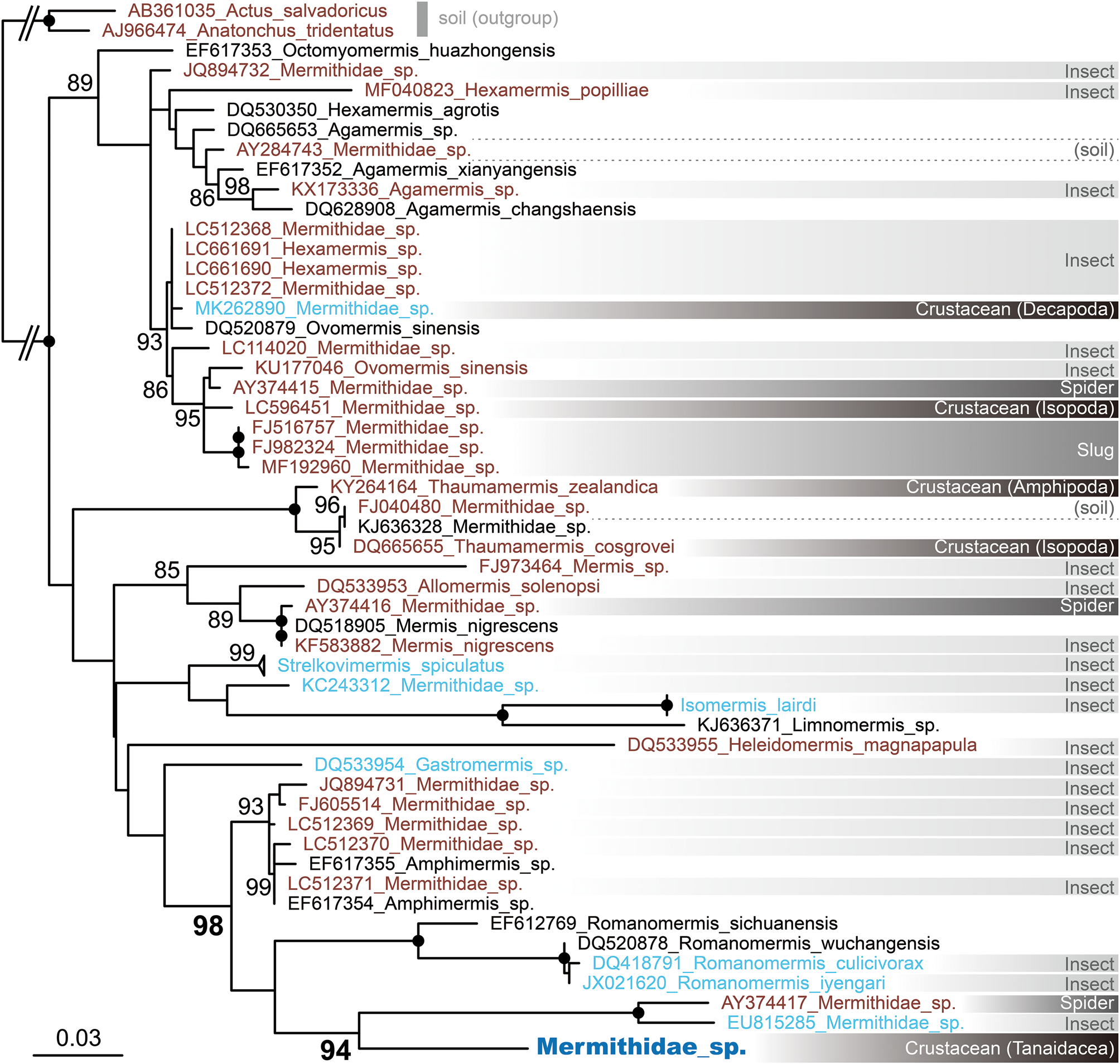
Fig. 2. ML tree for 18S sequences (dataset length 687 bp), including the sequence from Mermithidae sp. parasitic in Zeuxo sp. For the mermithid taxa, font colour indicates the environment where the host and/or the nematode individual used for sequence determination was collected: black, no data; brown, terrestrial; light blue, freshwater (includes host species, such as mosquitos, having an aquatic phase in their life cycle); dark blue, marine. Information on the host group is presented in the right-hand column, with different groups differentiated by different shades of grey. Numbers near nodes are ultrafast bootstrap values >80%; black circles indicate 100% ultrafast bootstrap support. Scale bar indicates branch length in substitutions per site.
This study provides the third example of mermithids from the marine environment and shows that tanaidaceans can serve as mermithid hosts. Knowledge of tanaidaceans as nematode hosts is currently somewhat limited. Only a few studies (Gardiner, Reference Gardiner1975; Tchesunov & Rozenberg, Reference Tchesunov and Spiridorov2011; Jakiel et al., Reference Jakiel, Palero and Błażewicz2019) have reported tanaidaceans as hosts for nematodes; Martell & McClelland (Reference Martell and McClelland1995) suggested that the anisakid nematode Pseudoterranova decipiens (Krabbe, 1878) uses tanaidaceans as an intermediate host. Except for the benthimermithid nematode Trophomera granovitchi Tchesunov & Rozenberg, 2011, none of the nematodes parasitic in tanaidaceans have been described morphologically. Future surveys for parasites in the small crustacean group Tanaidacea may provide additional examples of mermithid infections as well as those from other, previously unreported nematode groups.
Supplementary material
To view supplementary material for this article, please visit https://doi.org/10.1017/S0022149X22000256
Acknowledgements
We thank Koichi Morita, Masataka Kurosawa, Asako Matsumoto, Masato Hirose and Masanori Okanishi for their help in sampling; the staff of the International Coastal Research Center (Atmosphere and Ocean Research Institute, The University of Tokyo) for providing laboratory facilities; and Matthew H. Dick for reviewing the manuscript and editing our English.
Financial support
This study was supported in part by a KAKENHI grant (D.S., grant number JP21K06299) from the Japan Society for the Promotion of Science.
Conflicts of interest
None.
Ethical standards
The authors assert that all procedures contributing to this work comply with the ethical standards of the relevant national and institutional guides on the care and use of laboratory animals.





