SHEA Spring 2024 Abstracts
Poster Presentation - Poster Presentation
Water Management
Chlorine levels and a Legionella outbreak
- Sean Wu, Jyoti Somani, Hwang Ching Chan, Nazira Fauzi
-
- Published online by Cambridge University Press:
- 16 September 2024, p. s159
-
- Article
-
- You have access Access
- Open access
- Export citation
-
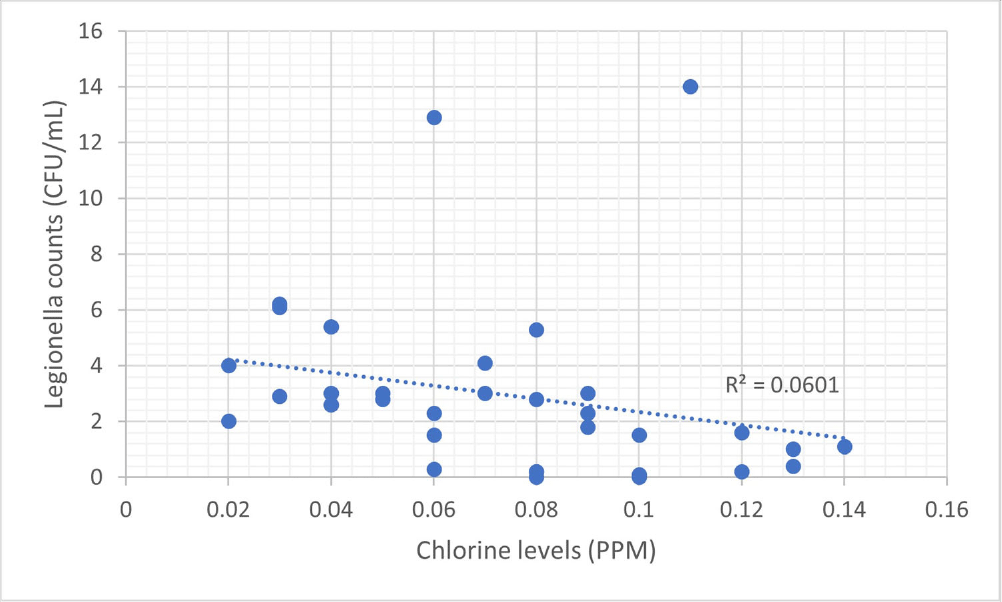
Background: Legionella, first identified in the 1970s, is increasingly recognized as an opportunistic pathogen in healthcare facilities.1 The National University Hospital (NUH) is a large quaternary level academic hospital with 1,200 beds located in equatorial Singapore. Its first building, completed in 1985, still serves patients today. In 2022, the infection prevention team (IPT) was informed of two cases of nosocomial legionella, which sparked the start of an extensive investigation consisting of water quality testing of multiple sources, case finding, and formation of a water management committee. Methods: 250mL water samples were collected and cultured by an external vendor using direct membrane filtration followed by plating on selective agar according to ISO 11731 standards. Bacterial colonies were then identified, quantified and speciated. At the same time, NUH’s facility management measured chlorine levels using a portable colorimeter. Results: In total, we cultured 34 samples taken from sinks, shower heads, potable water dispensers. 91.2% of samples were positive for legionella. Of the 91.2%, 35.3% of sites grew Legionella pneumophila serogroup 1 while 79.4% of sites grew Legionella pneumophila serogroup 2-15. We attributed our high rates of legionella positivity to an aging plumbing system and Singapore’s high humidity and temperatures. In addition, the maximal temperature of our hot water is only 48-50 °C. Although chlorine levels were generally low, they were still within the local recommendation of less than 2ppm (Singapore does not have guidance on minimum chlorine levels). We found no statistically significant correlation between the number of legionella colony forming units (CFUs) and chlorine levels (ranging between 0.02 to 0.14ppm). This supports the United States Environmental Protection Agency’s recommendation, as well as the findings from in vitro and in vivo studies, for a minimal chlorine of 0.2 PPM at the taps for acute care hospitals.2,3However, these levels may be inadequate in the presence of acanthamoeba or a high biofilm load within water systems.4,5 Conclusion: Hospital water management programs should require a minimal level of chlorine at hospital taps and at levels above those recommended by public water systems, in order to control legionella growth. In addition, the formation of a hospital water management committee is essential to improve hospital water quality and put mitigation measures in place. References 1. Phin, N. et al. Epidemiology and clinical management of Legionnaires’ disease. Lancet Infect. Dis. 14, 1011–1021 (2014). 2. Marchesi, I. et al. Monochloramine and chlorine dioxide for controlling Legionella pneumophila contamination: biocide levels and disinfection by-product formation in hospital water networks. J. Water Health11, 738–747 (2013). 3. Cervero-Aragó, S., Rodríguez-Martínez, S., Puertas-Bennasar, A. & Araujo, R. M. Effect of Common Drinking Water Disinfectants, Chlorine and Heat, on Free Legionella and AmoebaeAssociated Legionella. PloS One 10, e0134726 (2015). 4. Kessler, M. A., Osman, F., Marx, J., Pop-Vicas, A. & Safdar, N. Hospital-acquired Legionella pneumonia outbreak at an academic medical center: Lessons learned. Am. J. Infect. Control 49, 1014–1020 (2021).
Technology
An “Epic” Journey to Improve Antimicrobial Stewardship
- Lindsay Smith, John Ahern, Lisa Lapin, Thyleen Tenney
-
- Published online by Cambridge University Press:
- 16 September 2024, pp. s157-s159
-
- Article
-
- You have access Access
- Open access
- Export citation
-
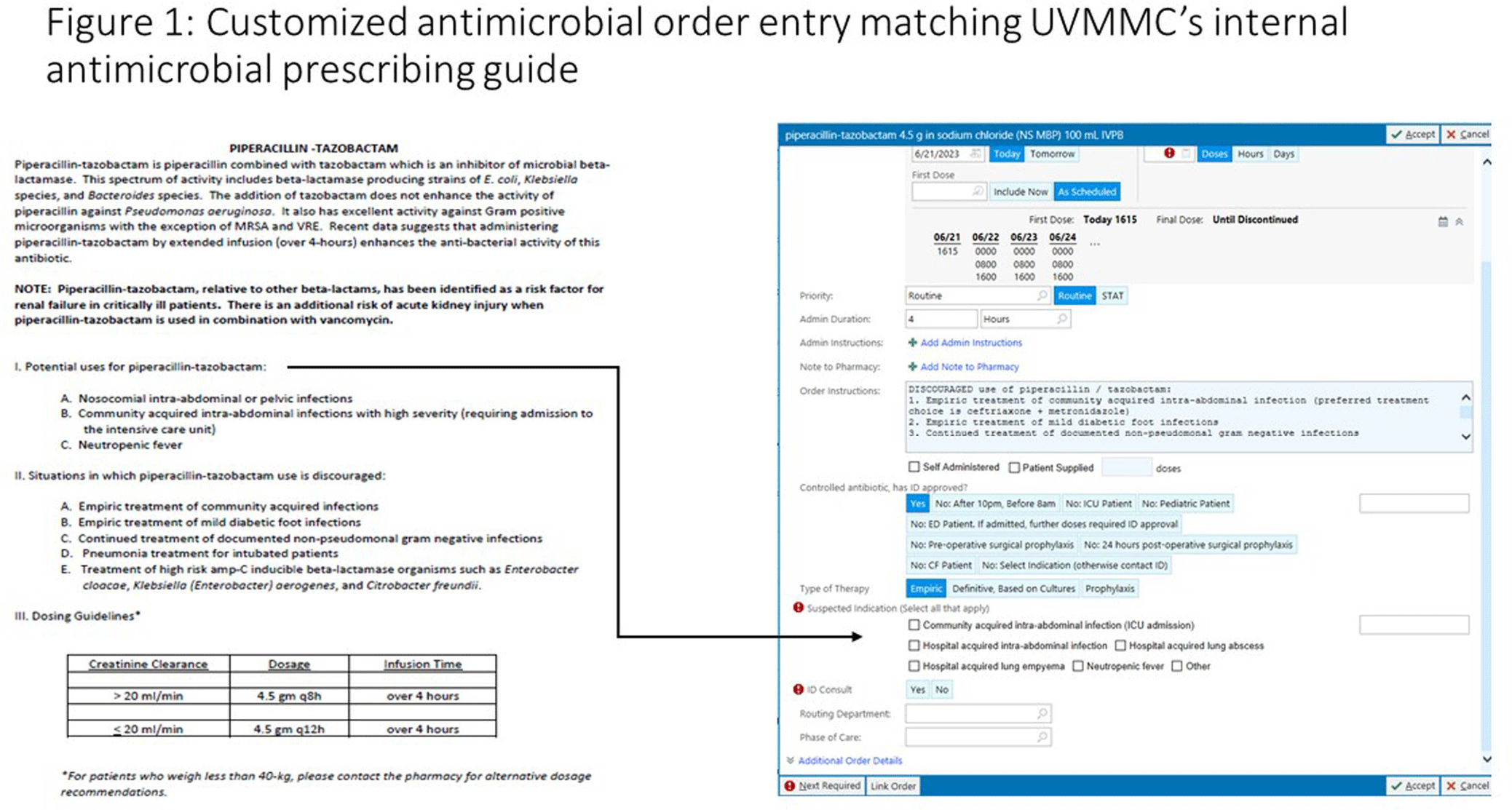
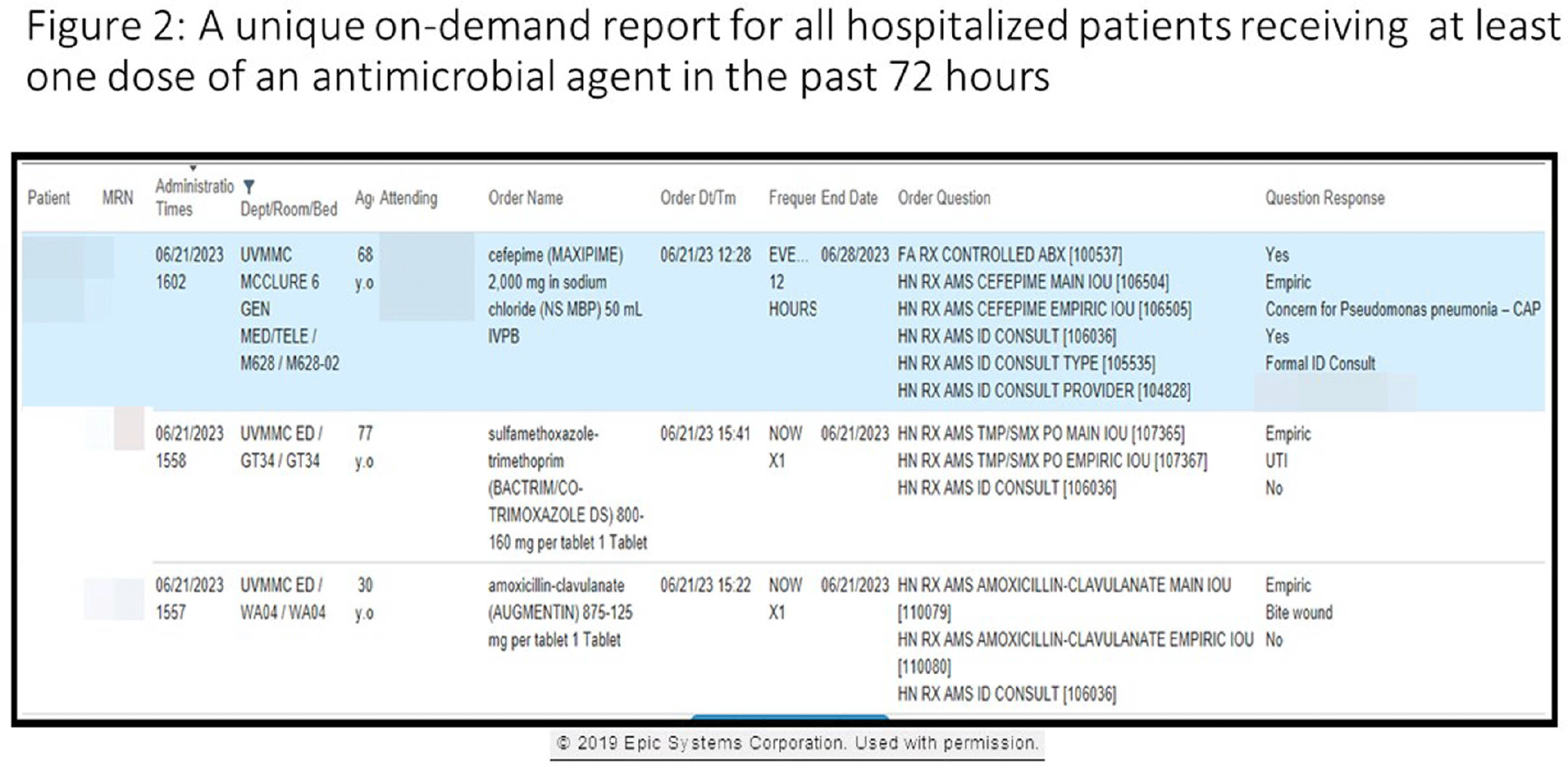
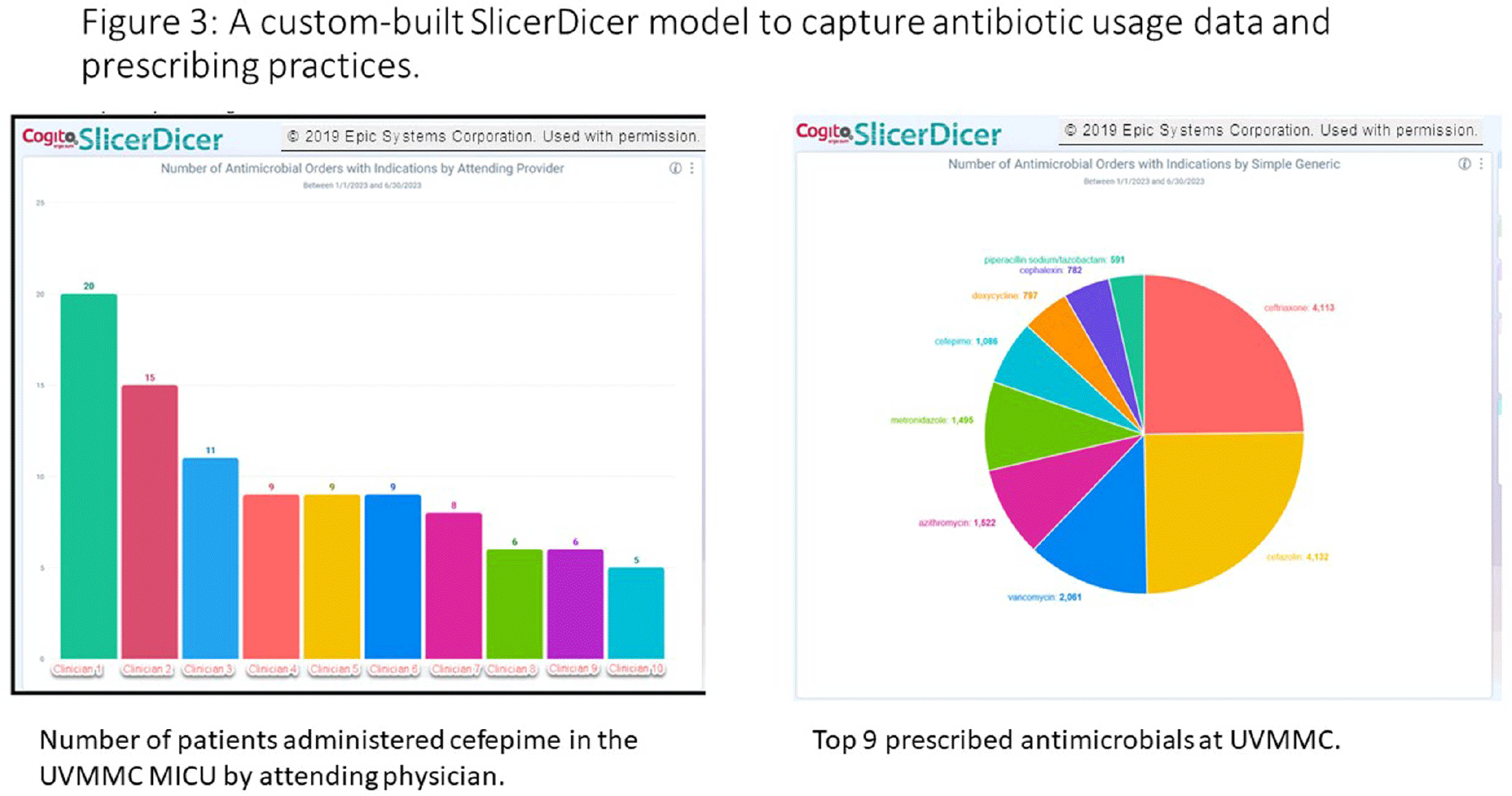
Background: Antimicrobial stewardship programs rely heavily on the electronic medical record (EMR) to carry out daily activities, make interventions, optimize patient care, and collect data. In 2019 the University of Vermont Medical Center transitioned from using a third party platform to the Epic (Verona, WI, www.epic.com) Bugsy module for antimicrobial stewardship. Method: We have spent the past 4 years optimizing the Epic foundation to match our institutional antimicrobial prescribing guidelines, susceptibility patterns, and build reports to extract actionable data. Result: During the build process, we readily identified three areas needed for customization: (1) Empiric, definitive, and prophylactic indications of use for all antimicrobials based on our hospital’s internally published books “Guide to Antimicrobial Therapy for Adults” and “Guide to Antimicrobial Therapy for Pediatrics” (figure 1); (2) An on-demand report to capture all patients with new administrations of antimicrobials in the preceding 72 hours, that includes ordering clinician, stop date of therapy, and indication (figure 2); and (3) A unique, custom-built slicer-dicer report to capture high-level data on how each antimicrobial is being prescribed by indication, dose, route of administration, ordering clinician, attending physician, and department (figure 3). Conclusion: We have built a system where we can readily identify patients that are receiving antimicrobials both within and outside of institutional guidelines and know the ordering clinician to contact to provide in-the-moment feedback. We can also collect retrospective data to know which antimicrobial agents were prescribed for all infectious syndromes. These three institutional customizations have provided invaluable information to improve patient care.
Surveillance
From Swiffers to Solutions: The Impact of Environmental Sampling in the Veterinary Medical Center at OSU
- Christy King, Thomas Wittum, Dixie Mollenkopf, Dubraska Diaz-Campos, Joany Van Balen
-
- Published online by Cambridge University Press:
- 16 September 2024, p. s157
-
- Article
-
- You have access Access
- Open access
- Export citation
-
Background: In 2018, the Ohio State University College of Veterinary Medicine (OSU CVM) implemented an Antimicrobial Stewardship Program, central to which was the integration of an environmental surveillance (ES) program. The ES focuses on pathogens recognized as urgent threats to public health by the Centers for Disease Control and Prevention. The pathogens currently targeted include carbapenemase-producing Enterobacterales (CPE), Salmonella spp., methicillin-resistant Staphylococcus spp. (MRSs), vancomycin resistant Enterococcus spp., and enrofloxacin resistant Pseudomonas aeruginosa. Identification of these pathogens allows the hospital to be aware of the local environmental microflora which can act as a sentinel for disease in the hospital, potentially causing healthcare associated infections. Therefore, the objective of this program is to identify resistant bacterial pathogens, characterize their resistance profiles, analyze prevalence patterns, and initiate infection control interventions where needed in the OSU VMC. Method: From January 2018 through December 2023, a total of 5449 samples were collected from approximately 86 locations across the OSU VMC encompassing the small animal, equine, and farm animal sections. A majority (64%, n=3561) of samples were collected from the small animal hospital, with the farm animal section contributing 1055 samples and the equine section 899. Areas sampled were frequented by both humans and animals, as well as surfaces exclusively touched by humans. Samples were collected using Swiffers® and processed through selective culture media. Result: Approximately half (52%, n=2890) of the samples collected represented human-touch only surfaces. A total of 3794 bacterial isolates were recovered, with an overall low prevalence for all targeted pathogens. Prevalence of CPE was 2% (n=103), with Enterobacter species being the most common. Recovery of MRSs was 8.5% (n=464) and Salmonella species was 1% (n=47). Conclusion: Through this initiative, the equine division of the OSU VMC collaborated with the antimicrobial stewardship team to enhance their Salmonella fecal and ES practices. In 2019, ES was critical in identifying persistent CPE and extended-spectrum cephalosporin-resistant Enterobacteriaceae in the ICU and surrounding areas of the small animal hospital. Effective measures were taken to halt the spread of ESC among patients and eliminate CPE in the environment. With the discovery of a new CPE in early 2023 in the small animal ICU and nearby areas, the program initiated targeted ES and cleaning and disinfection protocols, to identify contaminated areas and control disease transmission. These efforts have increased patient safety, health, and well-being, demonstrating how ES can be an important tool for infection control and prevention in veterinary settings.
Carbapenemase-Producing Enterobacteriaceae detected in a Large Canadian Tertiary Care Hospital: Five-year retrospective study
- Ashley James, Melisa Avaness, Victoria Williams, Lorraine Maze dit Mieusement, Heather Candon, Jerome Leis
-
- Published online by Cambridge University Press:
- 16 September 2024, p. s157
-
- Article
-
- You have access Access
- Open access
- Export citation
-
Background: The prevalence of carbapenemase-producing Enterobacteriaceae (CPE) is increasing worldwide. In Canada, where rates of healthcare-associated (HA) transmission of CPE remains relatively low, there is a need to share early experience of universal screening programs and risk factors for HA acquisition. Method: In 2018, universal screening was introduced throughout our large Canadian tertiary care hospital across, all critical care and oncology units. Additionally, risk-factor based screening was applied in all other inpatient units, with further targeted screening of roommate exposures or all inpatients on unit following identification of a single HA case. A retrospective cohort study was carried out on CPE cases detected between January 2018 and December 2023. We assessed the proportion of HA CPE cases, defined as CPE identified in patients with prior admission to our facility or after >72 hours after admission. HA cases were examined for relevant risk factors, including known roommate with CPE, the presence of other CPE on the unit, exposure to outbreak units, prior travel history, travel by a family member, and antibiotic exposure within the past 90 days. Result: A total of 150 CPE cases were identified, with 66 (44%) classified as HA. Among these HA cases, 14 (21%) were associated with presence of known case on the unit. The remaining 52 (79%) represented sporadic nosocomial cases without a known exposure or further transmission on the unit. Upon further retrospective review, 6 (9.2%) HA cases had documented travel history or exposure to a family member with recent travel to China, India, Sri Lanka, or the United States within the past year. Nearly all HA cases (62, 95.4%) had antibiotic exposure within 90 days of CPE detection; specifically, 47 (72.3%) received beta-lactams, 42 (64.6%) cephalosporin, 25 (38.5%) glycopeptide, 20 (30.8%) carbapenem, and 8 (12.3%) macrolide. Conclusion: HA CPE acquisition identified during the first 5-years of universal screening were mostly sporadic and not associated with known exposures or other risk factors. Receipt of prior antibiotics was present in nearly all cases.
Organism-specific Trends in Carbapenem-resistant Enterobacterales Infections in a Cohort of Hospitalized Patients, 2012–2022
- Mohammed Khan, Hannah Wolford, Natalie McCarthy, Babatunde Olubajo, Jonathan Bishop, James Baggs, Joseph Lutgring, Sujan Reddy, Maroya Walters
-
- Published online by Cambridge University Press:
- 16 September 2024, p. s156
-
- Article
-
- You have access Access
- Open access
- Export citation
-
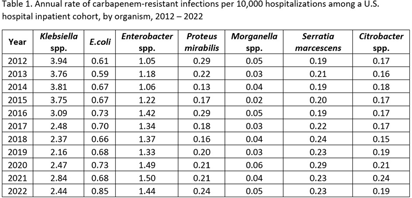
Background: Carbapenem-resistant Enterobacterales (CRE) infections are an urgent public health threat. An estimated 12,700 CRE (including E. coli, Klebsiella spp., and Enterobacter spp.) infections occurred in the United States in 2020. While the estimated incidence of CRE infections has been relatively stable between 2012 and 2020, organism-specific trends, including those for organisms not typically included in CRE surveillance definitions, have not been described. We estimated the annual rate of carbapenem-resistant Enterobacterales infections, disaggregated by organism, from 2012 to 2022. Methods: Data on inpatient hospitalizations from a dynamic cohort of short-term acute care hospitals reporting microbiology data between 2012 and 2022 were obtained from the PINC AI Database and the BD Insights Research Database. We included patients with clinical isolates of E. coli, Enterobacter spp., Klebsiella spp., Citrobacter spp., Serratia marcescens, Proteus mirabilis, and Morganella spp. and sufficient susceptibility results to identify carbapenem resistance. We limited our analysis to incident isolates, defined as a patient’s first isolate of a given organism and carbapenem resistance phenotype in a 14-day period. We calculated the annual rate of carbapenem-resistant infections per 10,000 hospitalizations for each organism. Results: There were 3,018,792 incident isolates from 55.8 million hospitalizations included in the analysis. Overall, 31,226 incident carbapenem-resistant isolates were identified. The rate of carbapenem-resistant infections varied by organism and over time (Table 1). The rate of carbapenem-resistant Klebsiella spp. infections appeared to decline from 3.94 in 2012 to 2.44 infections per 10,000 hospitalizations in 2022. The rate of carbapenem-resistant Enterobacter spp. infections appeared to increase from 1.05 in 2012 to 1.44 infections per 10,000 hospitalizations in 2022. The rate of carbapenem-resistant E. coli infections also appeared to increase, from 0.61 in 2012 to 0.85 infections per 10,000 hospitalizations in 2022. Rates of carbapenem-resistant Proteus mirabilis, Morganella spp., Citrobacter spp., or Serratia marcescens infections were similar in 2022 compared to 2012. Conclusions: Disaggregating data by organism revealed heterogeneous trends, with apparent increases in rates of carbapenem-resistant Enterobacter spp. and E. coli infections and apparent decreases in rates of carbapenem-resistant Klebsiella spp. infections. Organism-specific CRE analyses may provide additional insight into CRE epidemiology.
Case validation of bloodstream infections with an antibiotic-resistant organism
- Jennifer Ellison, Blanda Chow, Andrea Howatt, Logan Armstrong, Ted Pfister, Zhe Lu, Kathryn Bush
-
- Published online by Cambridge University Press:
- 16 September 2024, p. s156
-
- Article
-
- You have access Access
- Open access
- Export citation
-
Background: Bloodstream infections (BSIs) are an important cause of morbidity and mortality in severely ill patients, contributing to increased length of hospital stay and higher cost of care. Alberta Health Services Infection Prevention and Control (IPC) conducts inpatient surveillance of new episodes of BSIs with methicillin-resistant Staphylococcus aureus (MRSA), vancomycin-resistant enterococci (VRE) or carbapenemase-producing organisms (CPO) in 112 acute care facilities. A case-finding process was undertaken to verify the accuracy of BSI data entry. Methods: All positive MRSA, VRE or CPO blood cultures in 2021 were linked to the Inpatient Discharge Abstract Database (DAD) and the National Ambulatory Care Reporting System (NACRS) to identify new cases during acute care admissions. The results were then compared to surveillance records captured by infection control professionals (ICPs). Cases with unmatched culture date and/or encounter date and cases not identified by ICPs were screened by the study team with final decision made by ICPs. Results were analyzed by ARO and by % increase in number of surveillance records. Results: The laboratory linkage identified 286 new cases. Comparing to surveillance records (n = 248) captured by ICPs, 137 (57.3%) had matching collection dates and encounter dates, 85 (35.6%) had close matches on collection dates and encounter dates, 17 (7.1%) records had either matching collection dates or encounter dates, and 1 (0.4%) record did not have any matches on dates. There were 46 records identified in the laboratory data that were not in the surveillance system and 8 records that were in the surveillance system but not matched to the laboratory data. After review, 22 Surveillance records had data entry errors (1 CPO BSI, 20 MRSA BSI, and 1 VRE BSI), and there were 14 BSI records found to be missing (13 MRSA BSI, 1 VRE BSI). This represents a 6% increase in MRSA BSI and a 3% increase in VRE BSI identified in 2021 and no increase in CPO BSI. Conclusions: A laboratory validation to determine if BSIs with an ARO were missed during routine IPC surveillance identified a small proportion of missed bloodstream infections. The most common reason for the miss was admission through the emergency department with multiple blood cultures collected during a single admission. These results will be shared with the Infection Control program to facilitate correct BSI capture.
epiXact-ONT: Long-read whole genome sequencing for rapid outbreak detection and comprehensive plasmid transmission analysis
- Emma Briars, Ian Herriott, Allison Brookhart, Talia Hollowell, Nicole Billings, Miriam Huntley, Mohamad Sater
-
- Published online by Cambridge University Press:
- 16 September 2024, p. s155
-
- Article
-
- You have access Access
- Open access
- Export citation
-
Background: Healthcare associated infections (HAIs) are a major contributor to patient morbidity and mortality. HAIs are increasingly important due to the rise of multidrug resistant pathogens which can lead to deadly nosocomial outbreaks. Traditional methods for investigating transmissions are slow, costly and have poor detection resolution. In addition, plasmid transmission which can horizontally transfer critical resistance and virulence genes is not part of routine infection control practice due to lack of comprehensive and cost-effective methods capable of identifying both pathogen and/or plasmid transmission. Here we demonstrate the utility of the Oxford Nanopore Technologies (ONT) platform for whole genome sequencing (WGS) based pathogen and plasmids transmission analysis. Methods: We developed a rapid end-to-end process that includes sample preparation, sequencing optimized for generating long-reads and bioinformatics workflow customized for error-prone ONT data. We use Flye to generate de novo assemblies and a secondary bioinformatics step to identify each circular sequence. Individual circular sequences with an Ori (≥1) are identified. For pathogen clonality analysis we perform a pairwise mapping-based chromosomal sequences comparison eliminating need for an external reference genome. Similarly, individual plasmids are separated and compared pairwise. We annotate both the circularized chromosomal and plasmid sequences for known resistance and virulence genes. Results: We performed ONT (and confirmatory Illumina) sequencing of the genomes of 20 bacterial isolates originating from 5 HAI investigations previously performed at Day Zero Diagnostics using epiXact®, our Illumina-based HAI sequencing and analysis lab service. ONT-based clonality determination had 100% agreement with the Illumina based pipeline. We also found that using the outbreak-specific assembled genomes instead of an external reference increased the SNP-calling resolution in the ONT pipeline. We also identified sets of clonal isolates with both identical plasmids and distinct plasmids; as well as sets of non-clonal isolates with identical plasmids and distinct plasmids. In one subset of 7 multi-species isolates, we identified 2-7 circularized plasmid sequences in each isolate, all harboring known resistance genes. 4 plasmids were found in multiple isolates, with each plasmid appearing in between 2 and 4 distinct isolates. Notably, blaNDM was identified in at least 1 plasmid in each isolate. Conclusion: We demonstrate the utility of ONT for comprehensive HAI investigations, establishing the potential to transform healthcare epidemiology with rapid outbreak determination covering pathogen and plasmid transmission in < 2 4 hours from sample receipt.
Comparative Analysis of Healthcare-associated Bloodstream Infections & CLABSI Surveillance in A Singaporean tertiary Hospital
- Shalvi Arora, Sheena Jin Min Ong, Pinhong Jin, Aung Myat Oo, May Kyawt Aung, Yak Weng Darius Chan, Yang Yong, Ian Wee Liang En, Xiang Ying Jean Sim, Lai Chee Lee, Moi Lin Ling, Indumathi Venkatachalam
-
- Published online by Cambridge University Press:
- 16 September 2024, pp. s154-s155
-
- Article
-
- You have access Access
- Open access
- Export citation
-
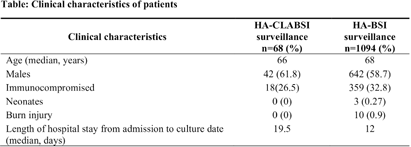
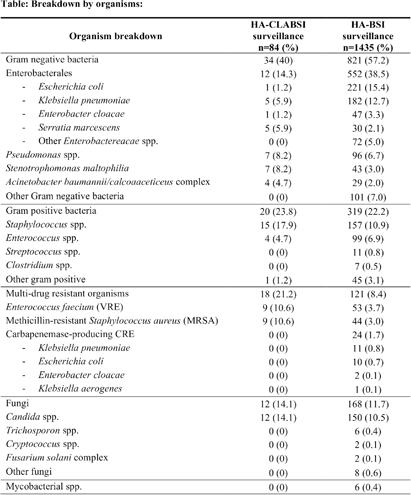
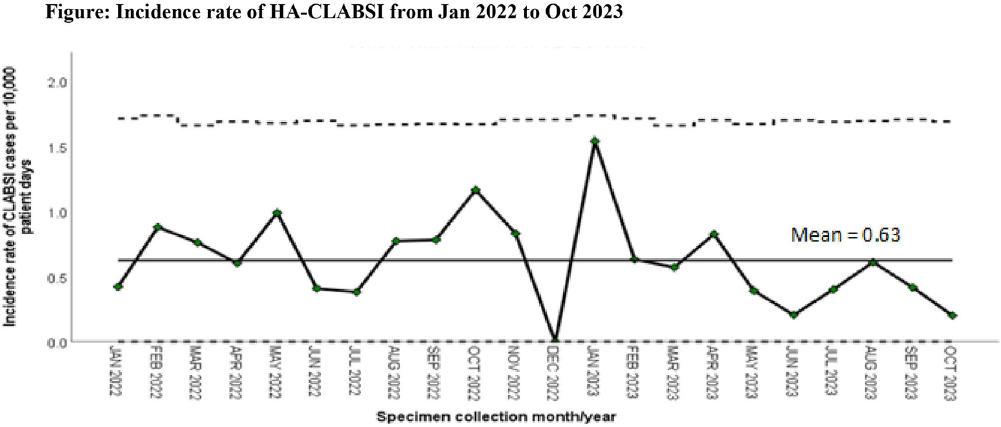
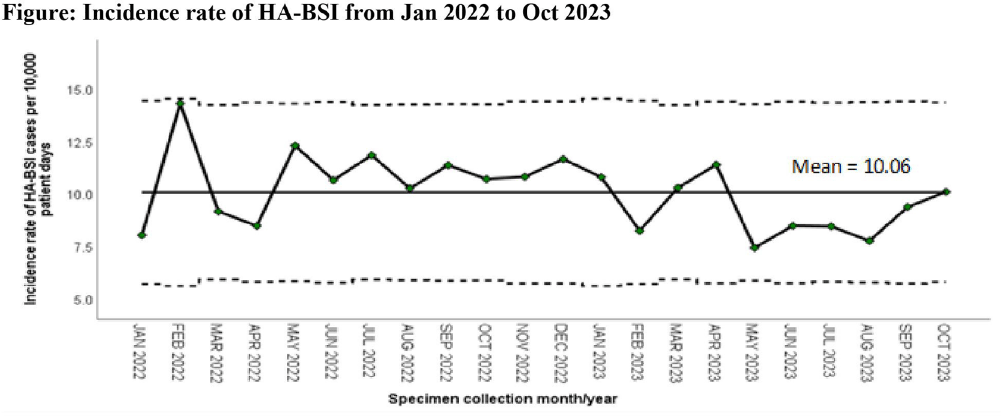
Background: Healthcare-associated central line associated bloodstream infection (HA-CLABSI) surveillance is important for monitoring healthcare-associated infections (HAIs) and evaluating effectiveness of infection prevention (IP) measures. However, implementing it is a laborious and time-consuming approach. Exclusive focus on central lines neglects HAI risk due to peripheral vascular catheters. This study aimed to assess whether HA-CLABSI incidence could be inferred from HA-bloodstream infection (BSI) trends and explore shift to HA-BSI surveillance. Methods: The study was performed in a Singaporean tertiary care hospital. Electronic medical records review was performed to determine whether positive blood cultures met Centers for Disease Control/National Health Safety Network (CDC/NHSN) definitions for HA-CLABSI and HA-BSI. Incident episodes of HA-BSI were included (excluding positive cultures repeated within 14 days). Incident organisms were explored to identify common causative pathogens (excluding same organisms isolated from cultures repeated within 14 days). CLABSI and BSI occurring ≥72hrs after admission were considered healthcare-associated. Patients under oncology or hematology service were considered immunocompromised. Incidence rates (IR) per 10,000 patient-days, patient characteristics and causative pathogens were compared between both indicators. Results: From January 2022 to October 2023, mean IR for HA-CLABSI was 0.63 (n=68) and for HA-BSI was 10.06 (n=1094). Median age of patients with HA-CLABSI was 66 years and HA-BSI was 68 years. HA-CLABSI and HA-BSI were more common in males (60.86% & 58.68%). Median duration between admission to HA-CLABSI was 20 days and to HA-BSI was 12 days. Median duration between central line insertion to HA-CLABSI was 16 days. Of 1094, 631 (57.7%) patients had vascular catheter(s) (i.e., IV cannula, port-a-cath, peripherally-inserted central catheter or central line) inserted at time of HA-BSI diagnosis, of whom 46 (7.3%) patients had CLABSI ±2days from positive blood culture. There was no significant correlation between monthly aggregate data from these indicators (Spearman’s correlation coefficient= 0.36, p-value=0.1). Predominant organisms causing HA-CLABSI and HA-BSI were gram negative bacteria (GNB, 40% & 57.21%), gram positive bacteria (24.71% & 22.23%), and fungi. Common GNB in CLABSI patients were Pseudomonas spp. and Stenotrophomonas maltophilia (8.24%), followed by Serratia marcescens and Klebsiella pneumoniae (5.88%). The frequent GNB in HA-BSI patients were Escherichia coli (15.4%), Klebsiella pneumonia (12.68%), and Pseudomonas spp. (6.69%). Common multi-drug resistant organisms were vancomycin-resistant Enterococcus faecium (10.59% & 3.69%) and methicillin-resistant Staphylococcus aureus (10.59% & 3.07%). Conclusion: HA-BSI did not correlate with HA-CLABSI. HA-BSI reflects heterogenous population outcomes. For utilization as surveillance indicator, further assessment on exclusion criteria is required to improve specificity.
Serratia marcescens Burden in a Neonatal Intensive Care Unit: Colonization Rate, Clinical Infections and Strain Relatedness
- Halima Dabaja Younis, Angelie Seguban, Andrea Morillo, Tony Mazzulli, Ming Lum, Jennie Johnstone
-
- Published online by Cambridge University Press:
- 16 September 2024, pp. s153-s154
-
- Article
-
- You have access Access
- Open access
- Export citation
-
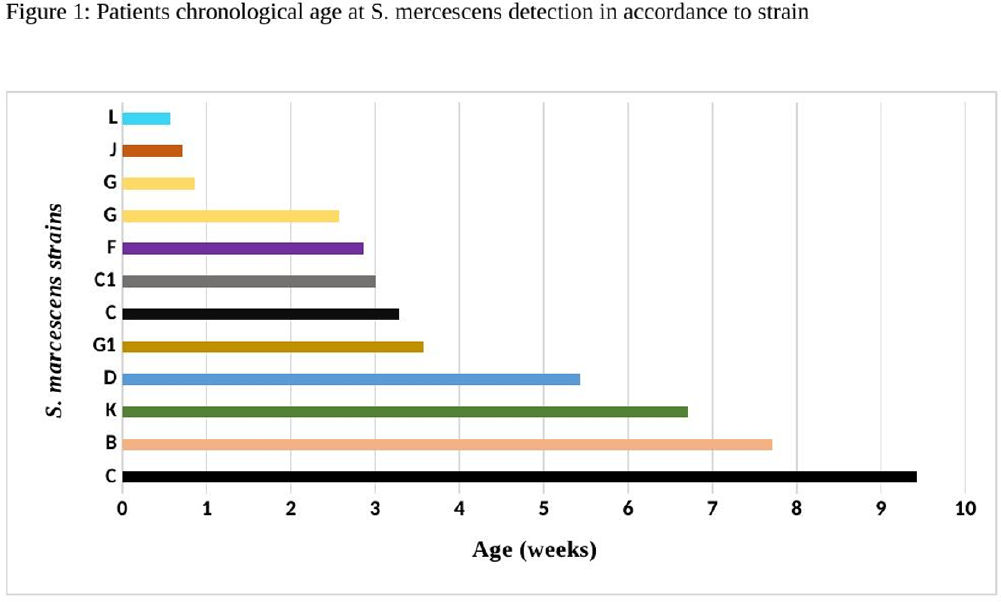
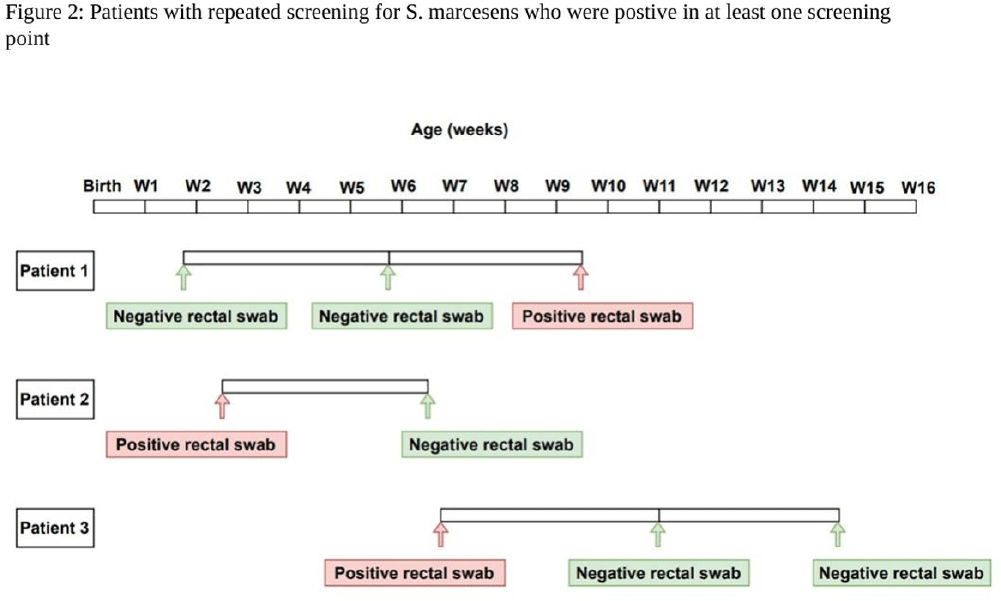
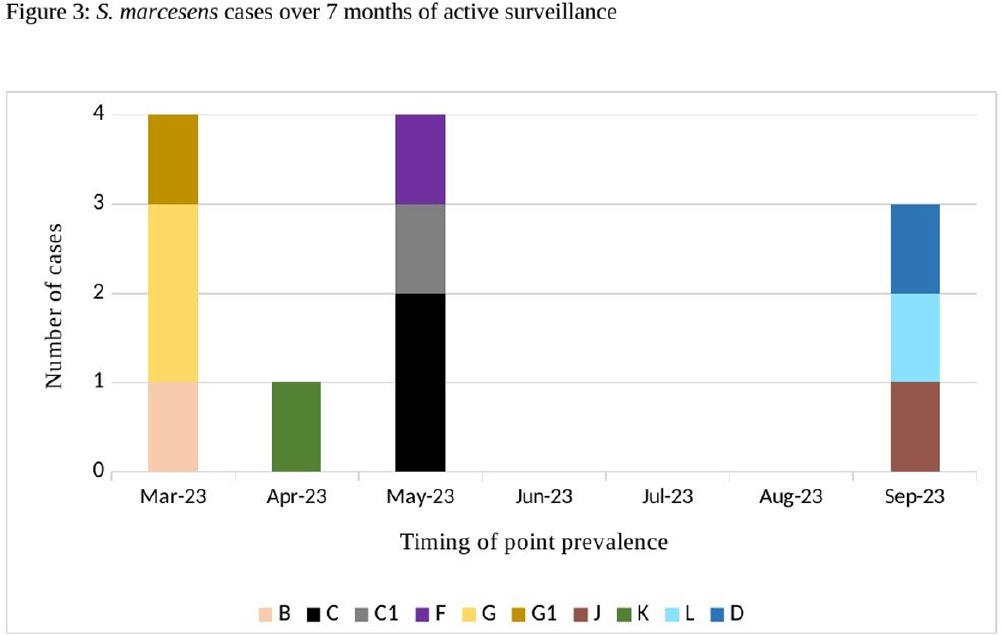
Background: Serratia marcescens (S. marcescens) is an environmentally associated organism known for causing healthcare associated infections and outbreaks in neonatal intensive care units (NICUs). The colonization or infection rates in NICU settings remain uncertain. This study aims to evaluate the rate of baseline colonization and clinical infection and relatedness of S. marcescens isolates. Methods: Prospective surveillance of rectal colonization and clinical infection of S. marcescens was conducted on patients admitted to the NICU at Mount Sinai Hospital in Toronto, Ontario, from March 1, 2023, to September 30, 2023. The NICU is a 57 bed unit with all private rooms. Monthly point prevalence assessments by rectal screening were performed, alongside active surveillance for clinical infections associated with S. marcesens. Isolates from screening or clinical samples underwent assessment for relatedness using pulse field gel electrophoresis (PFGE). Results: Over the 7 month study period, 12 different patients (5.4%) were colonized/infected with S. marcescens. Among these, 10 patients (4.5%) were identified through rectal screening (316 rectal swabs were collected from 224 patients) and two patients (0.9%) exhibited positive clinical specimens (urine and endotracheal aspirate) in association with pyelonephritis and ventilator-associated pneumonia, respectively. Of the two clinical cases, one case showed a negative preceding rectal swab and the other detected through a clinical sample before the point prevalence date. The age at which a positive S. marcescens swab or positive clinical specimen was identified ranged from 4 to 66 days (median=18 days, IQR 5-38.8) (Figure 1). Sixty seven infants had repeated screening. Three out of 67 (4.5%) were colonized with S. marcescens, the timing and sequence of positive and negative testing are presented in Figure 2. Females demonstrated a higher positivity rate compared to males [9.1% (9/99) vs 2.4% (3/125), p=0.04, respectively]. PFGE analysis of all 12 (100%) isolates revealed a polyclonal pattern. Most cases were detected from March to May in 9/12 cases (75%). Ten different strains were identified. Notably, two strains demonstrated clusters of two cases each, one during March and the other during May (Figure 3). No mortality was reported among the cases. Conclusions: The study highlights the polyclonal nature of S. marcesens and raises questions about the utility of point prevalence in anticipating clinical cases or patient-to-patient transmission, especially in patients with clinical infection where there were no preceding positive screening tests.
Variability of MDRO Reporting Across Tennessee Microbiology Laboratories
- Matthew Lokant, Christopher Wilson, Tom Talbot, Priscilla Pineda, Erin Hitchingham, Melphine Harriott, Raquel Villegas, Kaleb Wolfe, Milner Owens Staub
-
- Published online by Cambridge University Press:
- 16 September 2024, p. s153
-
- Article
-
- You have access Access
- Open access
- Export citation
-
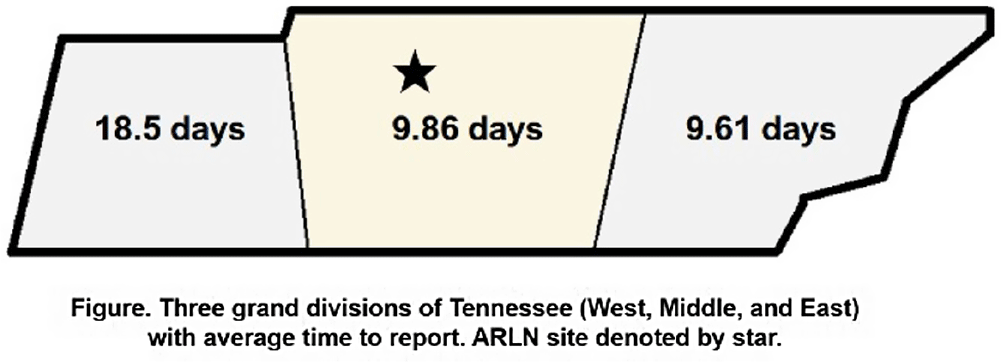
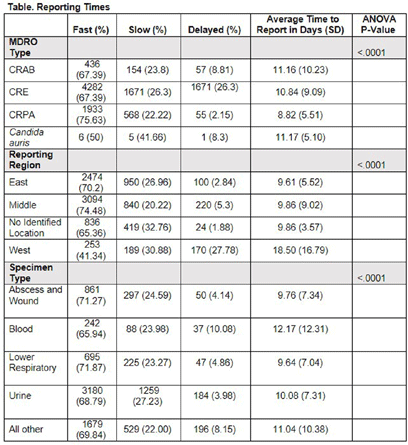
Background: Identification and timely reporting of multi-drug resistant organisms (MDROs) drives efficacy of infection prevention efforts. Data on MDRO reporting timeliness and inter-facility variability are limited. Facility-dependent variability in MDRO reporting across Tennessee was examined to identify opportunities for MDRO surveillance improvement. Methods: Data for reported Tennessee MDROs including carbapenem-resistant Enterobacterales (CRE), carbapenem-resistant Acinetobacter baumannii (CRAB), Carbapenem-resistant Pseudomonas aeruginosa (CRPA) and Candida auris, were obtained from the southeast regional Antibiotic Resistance Laboratory Network (ARLN) from 2018-2022, excluding screening and colonization specimens. Variance in days accrued from specimen collection to ARLN receipt was analyzed using one-way analysis of variance (ANOVA) with Tukey’s test (SAS 9.4). Facilities were categorized as fast (1-10 days), slow (11-20 days), or delayed (21-100 days) reporters. Results: There were 9,569 MDRO isolates reported. CRPA was reported faster than other MDROs (p < 0.001), while specimens from West Tennessee compared to other regions (p < 0.001) (Figure) and blood cultures compared to other specimens were reported more slowly (p < 0.001) (Table). There was no difference in reporting times for facilities using on-site microbiology laboratories versus reference laboratories (P = 0.062). Conclusion: MDRO reporting times varied across Tennessee by region, specimen, and organism. Future work to elucidate drivers of variability will consist of surveys and focused interviews with laboratory personnel to identify shared and unique barriers and opportunities for improvement.

Outpatient Antibiotic Consumption Trends in Belgium: A Comparative Analysis of Reimbursement and Sales Data, 2013-2022
- Elena Damian, Laura Bonacini, Moira Kelly, Boudewijn Catry, Lucy Catteau
-
- Published online by Cambridge University Press:
- 16 September 2024, pp. s152-s153
-
- Article
-
- You have access Access
- Open access
- Export citation
-
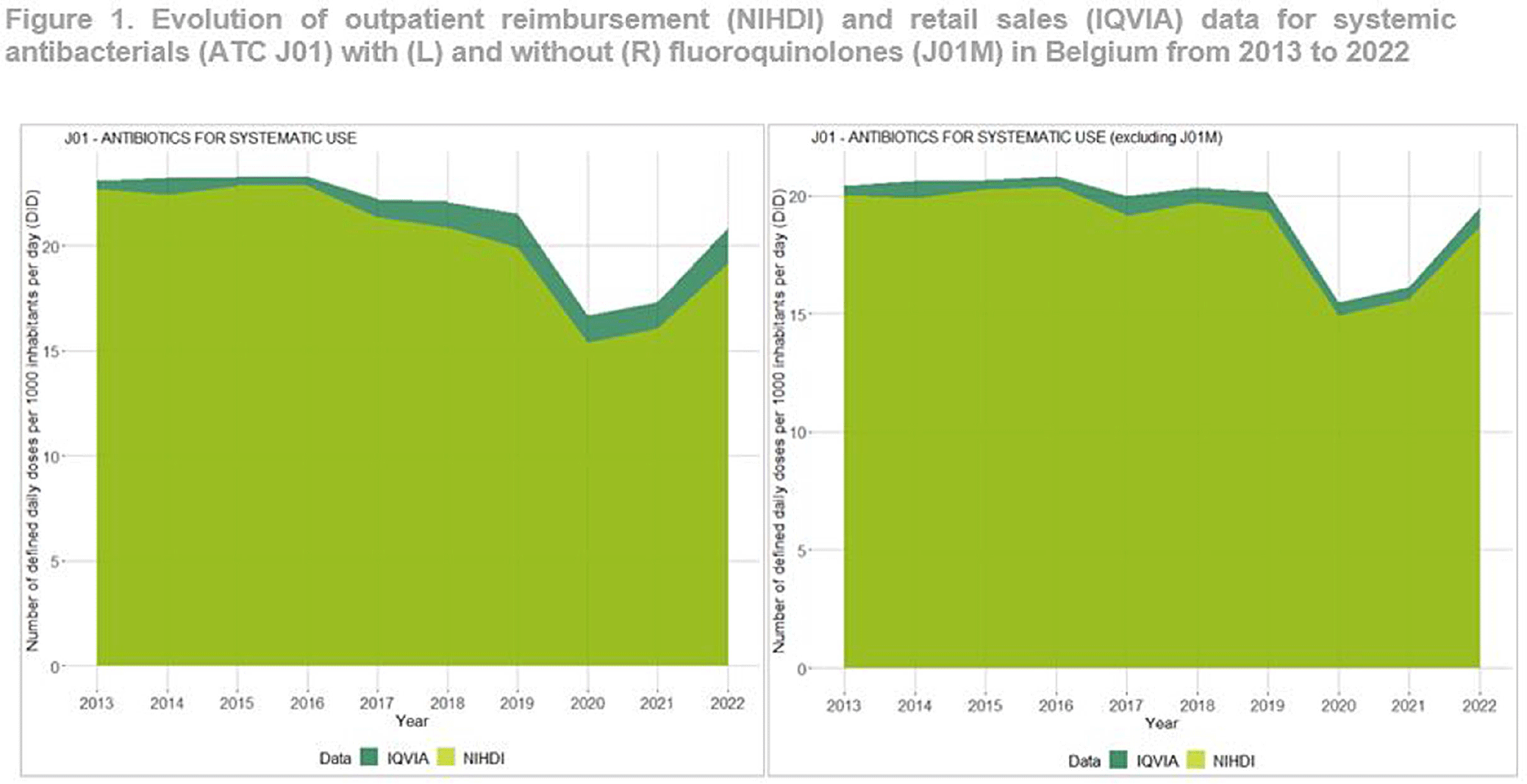
Background: Antimicrobial resistance (AMR) is a global public health concern, necessitating close and timely monitoring of antibiotic consumption (AMC). In Belgium, AMC surveillance traditionally relies on reimbursement data, excluding over-the-counter non-reimbursed or imported products and involving a time lag. This study investigates disparities in AMC between reimbursement data and retail data, providing insights into AMC variations. Additionally this study seeks to critically evaluate the validity and representativeness of the reimbursed data in accurately reflecting the true extent of AMC in the country. Method: Utilizing reimbursement data from the National Institute for Health and Disability Insurance (NIHDI) and retail data (IQVIA Sales data; www.iqvia.com) for systemic antibacterials (ATC Group J01), outpatient consumption was estimated for the period 2013-2022. Volume of antimicrobials was measured in Defined Daily Doses (DDDs - WHO ATC/DDD Index 2023), while population data were extracted from Eurostat. Relative differences (RDs) in DDDs per 1000 inhabitants per day (DID) were computed, and validated through correlation analysis (Pearson’s r) and Bland–Altman plots. Result: J01 antibacterial sales declined from 23.10 DID (2013) to 20.85 (2022). Non-linear decreases, notably during the Covid-19 pandemic (21.54 DID in 2019 to 16.69 in 2020), followed by a rebound to pre-pandemic quantities in 2022 were observed (Figure 1). Reimbursement NIHDI data slightly underestimated IQVIA sales, with RDs ranging from 2% (2013) to 9% (2022). Notable differences, especially in recent years were attributed to quinolone reimbursement criteria changes implemented by law in Belgium in 2018, reducing the reimbursed proportion from 99% (2017) to 35% (2022). ATC-3 level analysis revealed disparities in low-DID groups (J01B, J01E and J01G). Notably, a small proportion of amphenicols (J01B) were reimbursed ( < 1 0%), with a congestion relieving combination product of tiamphenicol (+ N-acetylcysteine; Fluimucil®) frequently bought and remaining unreimbursed. Overall and across ATC3 groups, the correlation between NIDHI and IQVIA estimates was almost perfect across years and the Bland–Altman plots showed high agreement. Conclusion: Reimbursement data are reliable for outpatient AMC monitoring with slightly lower estimates than retail data across most categories. The 2018 quinolone reimbursement criteria change highlights the necessity of incorporating retail data for accurate assessments in this specific category. The synergistic use of reimbursement and retail datasets is crucial for a comprehensive understanding of consumption patterns, supporting effective AMR mitigation strategies in Belgium.
An Improved Algorithm for the Detection of Ventilator Associated Pneumonia
- Dan Ding, Heidy Wang, Madeline DiLorenzo, Sarah Hochman, Corinne Thompson, Michael Phillips, Sherif Shoucri
-
- Published online by Cambridge University Press:
- 16 September 2024, pp. s151-s152
-
- Article
-
- You have access Access
- Open access
- Export citation
-
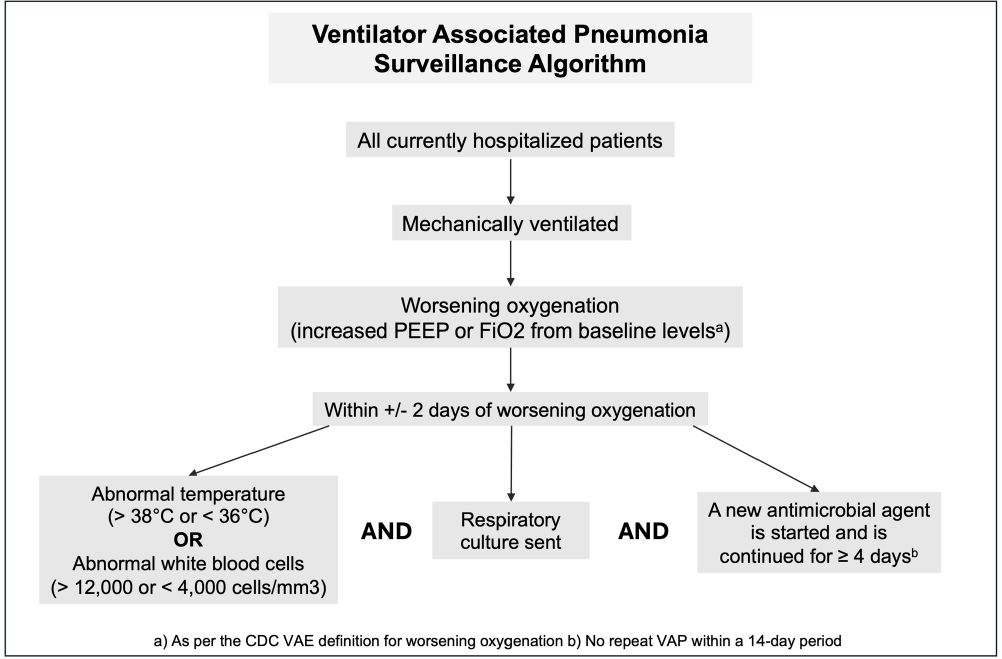
Background: Ventilator associated pneumonia (VAP) is associated with significant rates of morbidity and all-cause mortality. Active VAP surveillance can identify risk factors for which targeted preventive measures can be implemented. However, surveillance efforts are complicated by challenges associated with accurate VAP diagnosis. We aimed to improve the accuracy and automation of existing VAP diagnostic algorithms to better identify patients at risk. Methods: The study was conducted at NYU Langone Health from June 2022 through December 2023. We created a semi-automated VAP surveillance system using the Centers for Disease Control & Prevention (CDC) ventilator associated event (VAE) definition as a base framework (Figure 1). We modified this definition to include additional elements, such as having a sputum culture ordered within 48 hours of worsening oxygen status, regardless of culture result. Using this algorithm—followed by manual clinician reviews—we retrospectively assessed possible VAP cases to determine the ability of our surveillance system to correctly identify VAP. Results: Of the 123 possible VAP cases identified through our automated system, 75 (61%) were correctly diagnosed as VAP after clinical review. This reflects a rate of 1.5 infections per 1000 ventilation days across the system and 1.85 infections per 100 patients ventilated for greater than 2 days. Of the 48 remaining patients without VAP after clinical review, 25% (n=12) were characterized as having hospital-acquired pneumonia, 21% (n=10) as acute respiratory distress syndrome or infection at another site and 10% (n=5) as pulmonary embolism/infarction. Among all patients identified through this automated system (VAP and non-VAP), 53% experienced in-hospital death. Discussion: Our automated VAP surveillance algorithm identified 123 cases of potential VAP, 61% of which were consistent with a clinical diagnosis of VAP upon manual chart review. Our VAP rate of 1.5 infections per 1000 ventilation days was similar to published rates at other North American hospital systems. The high in-hospital mortality rate among these patients highlights the need for improved surveillance systems and earlier interventions to reduce the risk of VAP. There are several limitations to the CDC’s VAE definition, including its requirement of a positive microbiologic culture and focus on sputum quality. This potentially misses cases of culture-negative VAP in patients receiving antibiotics prior to sputum collection. Our goal is to continue to validate and improve our algorithm’s ability to correctly identify patients with clinical VAP, so that targeted prevention efforts can be focused upon the patients with the highest risk for poor outcomes.
Disclosure: Madeline DiLorenzo: Stocks - Abbvie, Amgen Inc., Becton Dickinson, Biogen Inc., Bristol Myers and Squibb, CVS Health, Davita Inc., Elevance Health, Gilead, Henry Schein, Hologic Inc., Humana Inc., Jazz Pharmaceuticals, Laboratory Corp, Merck and Co., Quest Diagnostics, ResMed Inc., Teladoc Health, Vertex Pharmaceuticals, West Pharmaceuticals
Impact of Point Prevalence Surveillance on Transmission of Candida auris Within an Acute Care Health System
- Joan Ivaska, Milena Walker, Claire Jai, Marjorie Bessel
-
- Published online by Cambridge University Press:
- 16 September 2024, p. s151
-
- Article
-
- You have access Access
- Open access
- Export citation
-
Background: Patients infected or colonized with Candida auris can serve as a transmission source for other patients. Screening patients for Candida auris colonization allows facilities to implement infection prevention and control measures and minimize the risk of transmission. The Centers for Disease Control and Prevention (CDC) recommends healthcare facilities perform three types of screening; admission screening in patients with specific risks, close contact screening of patients who overlap a confirmed positive case for 3 or more days or point prevalence surveillance if there is evidence of ongoing transmission within the facility. The CDC further recommends that patients being screened for Candida auris be maintained in transmission-based precautions while awaiting Results: In 2022-2023 there was ongoing transmission of Candida auris occurring in a community served by a large multi-state healthcare system. Close contact and point prevalence surveillance screening for both acute and non-acute healthcare facilities were implemented by the local health jurisdiction. Methods: A composite swab of the bilateral axilla and groin was used to screen close contacts of patients confirmed to be infected or colonized with Candida auris. Close contact was defined as having been on the same unit as the positive patient for 3 or more days while the patient was not in transmission-based precautions. Point prevalence surveillance was performed on all patients currently housed on units where close contact screen-positive patients resided. Potentially exposed patients who had been discharged were not screened. Patients were placed in contact transmission-based precautions until results were received. In 1657 patients in six acute care facilities were identified for Candida auris screening. 161 patients refused or were unable to be screened. Of the 1496 patients screened, 40 screened positive, demonstrating a 2.67% secondary attack rate. Of the 40 screen-positive patients, 5 were identified through point prevalence and 35 through close contact screening. Conclusion: Performing point prevalence surveillance in acute care facilities is operationally challenging and costly with little benefit in the prevention of Candida auris transmission. More robust collection and reporting of screening data is needed to inform surveillance protocols and prevention strategies specific to different healthcare settings. Limitations of this study include the lack of screening completion in discharged patients identified as close contact or point prevalence surveillance eligible. Additionally, some patients had a history of contact with healthcare facilities outside of this healthcare system, with unknown exposure risks or prevention strategies.
Antimicrobial Use in Belgian Acute Care Hospitals : Results of the 2022 ECDC Point Prevalence Survey
- Lucy Catteau, Katrien Latour, Morgan Pearcy, Boudewijn Catry
-
- Published online by Cambridge University Press:
- 16 September 2024, pp. s150-s151
-
- Article
-
- You have access Access
- Open access
- Export citation
-
Background: Point prevalence surveys (PPS) organized by the European Centre for Disease Prevention and Control (ECDC) play a crucial role in assessing healthcare-associated infections (HAIs) and antimicrobial use (AU) in European acute care hospitals. In 2017, a crude prevalence of 28.1% (95% CI 27.3-29.0%) of inpatients receiving at least one antimicrobial was recorded in Belgium (patients ≥65 years: 29.6% (95% CI 28.5-30.7%), < 6 5 years : 26.5% (95%CI 25.3-27.6%)) . Following the challenges posed by the COVID-19 pandemic, the 2022 ECDC-PPS aimed to reassess AU levels. Method: A cross-sectional study was conducted between September and November 2022 in 57 representative acute care Belgian hospital sites (35 mergers), following the ECDC-PPS protocol version 6.0. All patients present in surveyed wards at 8 a.m. on the PPS day and not discharged at that time were included. Infection prevention and control teams collected comprehensive data on hospitals, wards, and AU, including agents and indications. Results: Among the 10,142 included inpatients, 29.3% (95%CI 28.4-30.2) were receiving at least one antimicrobial (patients ≥65 years: 31.1% (95% CI 29.7-32.4%), < 6 5 years : 27.1% (95%CI 25.6-28.6%)). Intensive care units (56.3%), surgical (38.7%), and medical wards (33.1%) demonstrated the highest AU prevalence, while psychiatric wards exhibited the lowest (3.0%). A total of 3,549 antimicrobials were recorded, commonly prescribed for treating community-acquired infections (48.6%) and HAIs (30.3%, including 4.2% of long-term care facility acquired infections), as well as for surgical and medical prophylaxis (12.4 and 6.6%, respectively). Notably, only 22.7% of surgical prophylaxis courses (n=100/440) lasted more than one day. The top three most used antimicrobial agents consisted of amoxicillin in combination with a beta-lactamase inhibitor (J01CR02, 20.0%), cefazolin (J01DB04, 9.8%) and piperacillin in combination with a beta-lactamase inhibitor (J01CR05, 9.6%). The most frequently reported diagnoses for medical antimicrobial treatment were pneumonia (25.7%) and urinary tract infections (17.1%). The reason for AU was available in 80.0% of the medical notes. Conclusion: The 2022 PPS reveals an increased AU prevalence (+1.2%) in Belgian acute care hospitals, especially in patients over 65 years of age (+1.5%). This increase was less pronounced in younger patients (< 6 5y) (+0.6%). Future investigations are crucial to delve into prescription attitudes and modifiable practices, emphasizing the urgent need for robust antimicrobial stewardship programs in these healthcare settings.
Efficacy of Empiric Contact Precautions for Patients from High Risk Facilities
- Kavitha Prabaker, Dan Uslan, Annabelle De St. Maurice, Shaunte Walton, Vanessa Lewis, Anjali Bisht, Sebora Turay, Urvashi Parti, Ricardo Ison, Donna Wellbaum, Yvonne Mugford
-
- Published online by Cambridge University Press:
- 16 September 2024, p. s150
-
- Article
-
- You have access Access
- Open access
- Export citation
-
Background: Infection prevention surveillance revealed that patients admitted from two specific long term care facilities comprised the majority of multi-drug resistant organisms (MDRO) and scabies cases at our institution. Current practices include performing active surveillance for Candida auris and methicillin-resistant Staphylococcus aureus (MRSA) for specific high-risk patients, as surveillance for all MDROs and scabies is impractical. We therefore sought to create an admission screening process to efficiently identify patients from high-risk facilities (HRFs) and place them in pre-emptive contact precautions upon admission. Methods: Patients admitted from HRFs were identified on admission as part of the initial nursing assessment. For any positive responses, nursing received a Best Practice Advisory to place the patient in contact precautions and patient placement received an alert that the patient would require a private room. Infection Preventionists reviewed a report of all patients who screened positive and added a “High Risk Facility” banner to the chart. This banner remained for the duration of hospitalization and for every subsequent readmission and outpatient visit. We reviewed the electronic medical records of all patients with a HRF banner placed from March 8, 2023 to September 15, 2023 and abstracted data regarding the presence of scabies or any of the following MDROs before and after placement of the banner: C. auris, carbapenem-resistant enterobacterales (CRE), MRSA, vancomycin-resistant Enterococcus (VRE), carbapenem-resistant Acinetobacter, and MDR Pseudomonas. Results: Of the 93 patients who had a HRF banner added during the study period, 31 (33.33%) were already known to have MDRO colonization at the time of admission to our facility. Thirty-three of the remaining 62 patients (53.22%) without known MDRO colonization were subsequently found to have MDRO colonization/infection or scabies infestation that may have required contact precautions during their index admission or a subsequent admission. This included 14 patients with C. auris, 2 with CRE, 3 with MDR Pseudomonas, 12 with MRSA, 12 with carbapenem-resistant Acinetobacter, and 2 with VRE. Patients were admitted for a median of 9 days before their diagnosis, and 36 of the 93 patients (38.71%) were re-admitted to our hospital during the study period. Conclusion: We found that empiric contact precautions based solely on exposure to specific HRFs facilitated earlier isolation by a median of 9 days. This approach should be considered in acute care hospitals with a high proportion of admissions from HRFs, especially when active and passive surveillance for MDROs is limited.
Trends in Hospital Antibacterial Consumption in Belgium (2017-2021): Evaluating the Impact of the COVID-19 Pandemic
- Laura Bonacini, Elena Damian, Boudewijn Catry, Lucy Catteau
-
- Published online by Cambridge University Press:
- 16 September 2024, pp. s149-s150
-
- Article
-
- You have access Access
- Open access
- Export citation
-
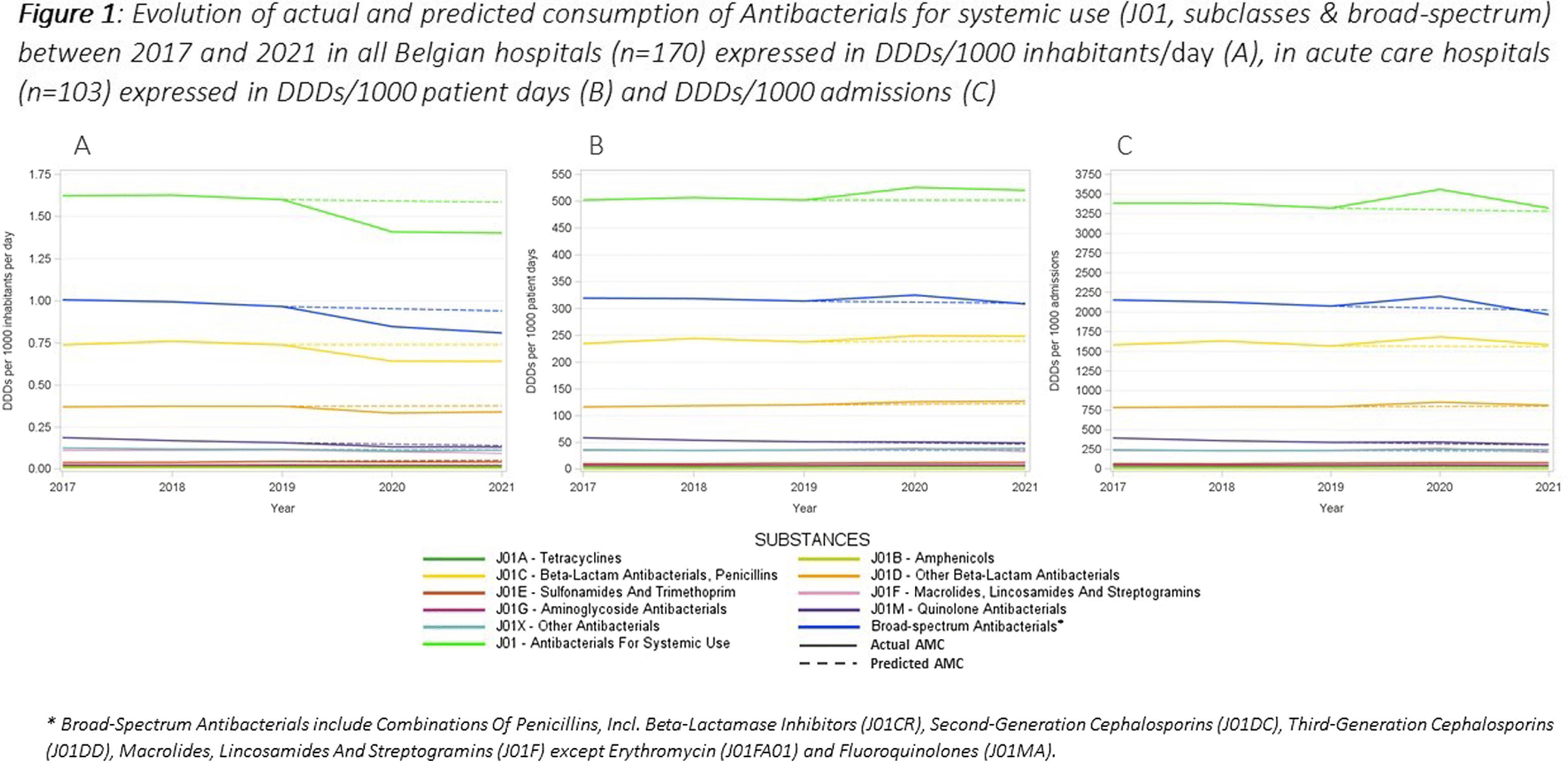
This study aimed to evaluate the impact of the COVID-19 pandemic on antimicrobial consumption (AMC) in Belgian hospitals from 2017 to 2021, using data from the European Surveillance of Antimicrobial Consumption Network (ESAC-Net) and the Belgian Hospitals Surveillance of Antimicrobial Consumption (BeH-SAC). Antimicrobial volume was quantified in Defined Daily Doses (DDDs), and AMC was expressed in DDDs/1000 inhabitants/day (DIDs), DDDs/1000 patient days and DDDs/1000 admissions. Linear regressions were employed to analyze 5-year trends for the ATC J01 group, at the ATC-3 level and for broad-spectrum antimicrobials. Broad-spectrum antibiotics included combinations of penicillins, incl. beta-lactamase inhibitors (J01CR), second-generation cephalosporins (J01DC), third-generation cephalosporins (J01DD), macrolides, lincosamides and streptogramins (J01F, excluding erythromycin J01FA01), and fluoroquinolones (J01MA). The compound annual growth rate (CAGR) calculated for the years preceding the pandemic was used to forecast 2020 and 2021 AMC, enabling a comparison with the actual use. Hospital AMC measured as DIDs decreased by 12% from 2019 to 2020. In contrast, when expressed as DDDs/1000 patient days and DDDs/1000 admissions, a 5% and 7% increase was observed, respectively. Antibacterials for systemic use (J01) showed a significant decrease over the 5 years only when expressed in DIDs. Notable trends included a negative trend for quinolone antibacterials (J01M) when expressed in the three incidence units, as for amphenicols (J01B) when using hospital denominators only. Positive trends were observed for sulfonamides and trimethoprim (J01E) using hospital denominators and for other beta-lactam antibacterials (J01D) with the ‘patient days’ denominator. While the consumption of all J01 antimicrobial subclasses deviated negatively from predicted use both in 2020 and 2021 when expressed in DIDs, positive deviations were recorded using hospital denominators, except for macrolides (J01F). The use of broad-spectrum antimicrobials showed a notable decrease between 2017 and 2021 when expressed in DIDs. However, when using hospital denominators, the observed use of broad-spectrum antimicrobials exceeded the forecasted values in 2020, to regress below the forecasted levels in 2021 (Figure 1). Contrary to results obtained using the widely applied country’s population as the denominator, a notable surge in AMC, particularly for broad-spectrum antimicrobials, was observed in 2020 when using hospital-specific denominators. This increase coincided with the onset of the COVID-19 crisis. These findings emphasize the need for a national hospital surveillance system that uses denominators that accurately represent the specific population being monitored. Implementing robust hospital-specific surveillance mechanisms would improve the precision of evaluations and facilitate targeted interventions aimed at optimizing antimicrobial utilization.
Comparison of COVID-19 Sentinel Surveillance and COVID-19 School Absentee Surveillance in Japan
- Masaki Tanabe, Shiho Ito, Yasuyuki Hara, Takahiro Ogura, Makoto Ichikawa, Miwa Fukuta, Yoshito Iwade
-
- Published online by Cambridge University Press:
- 16 September 2024, pp. s148-s149
-
- Article
-
- You have access Access
- Open access
- Export citation
-
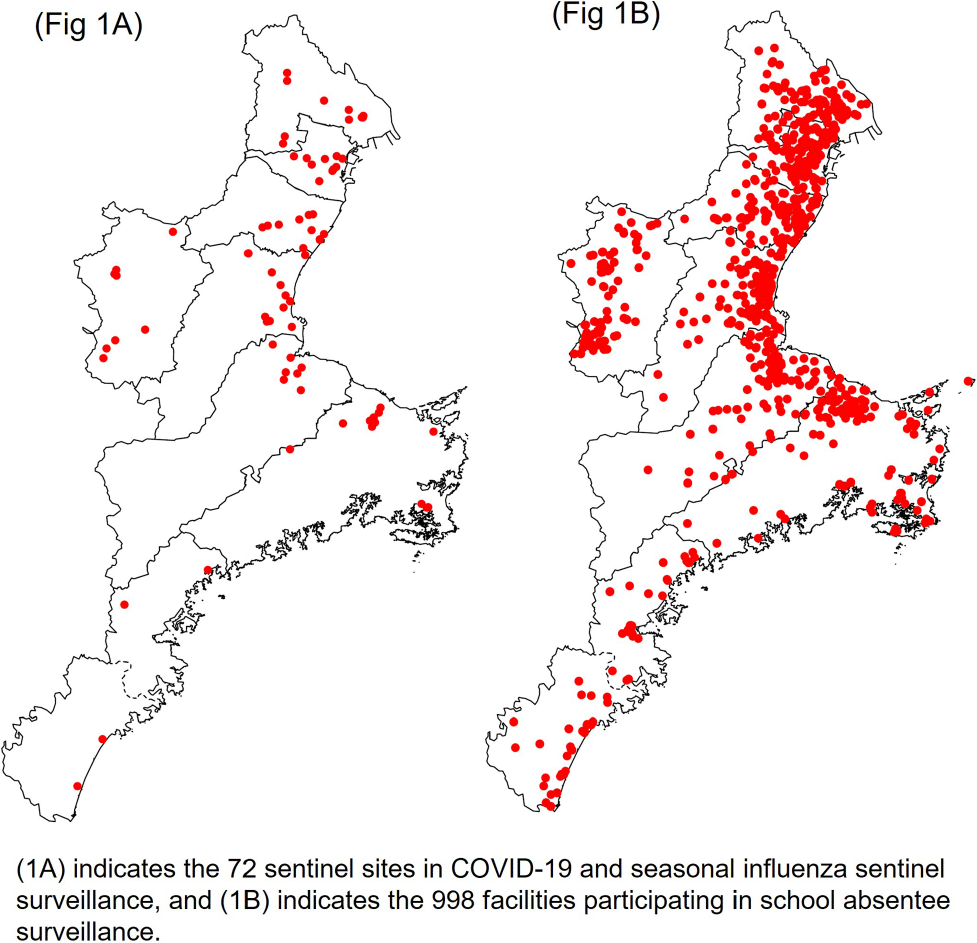
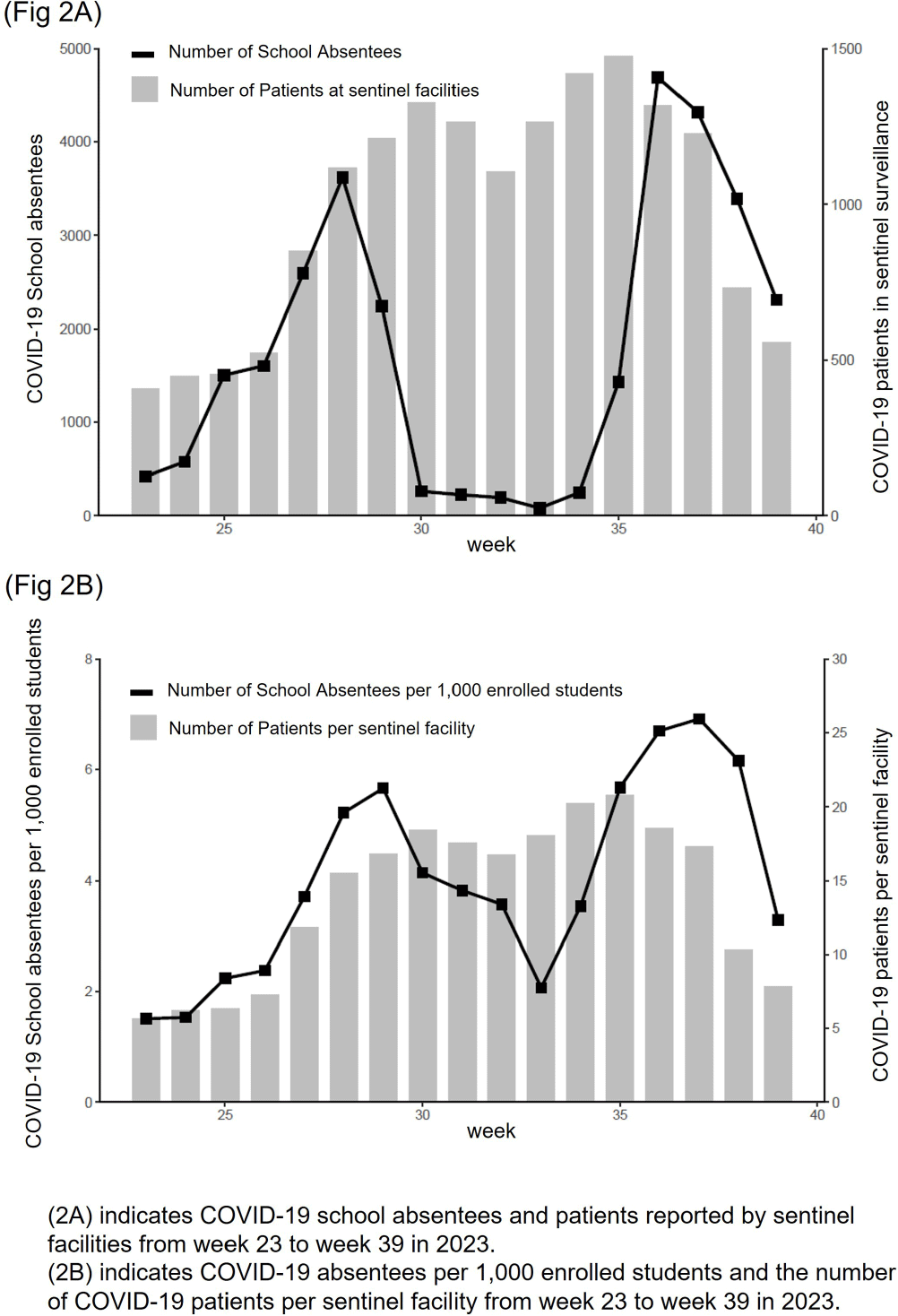
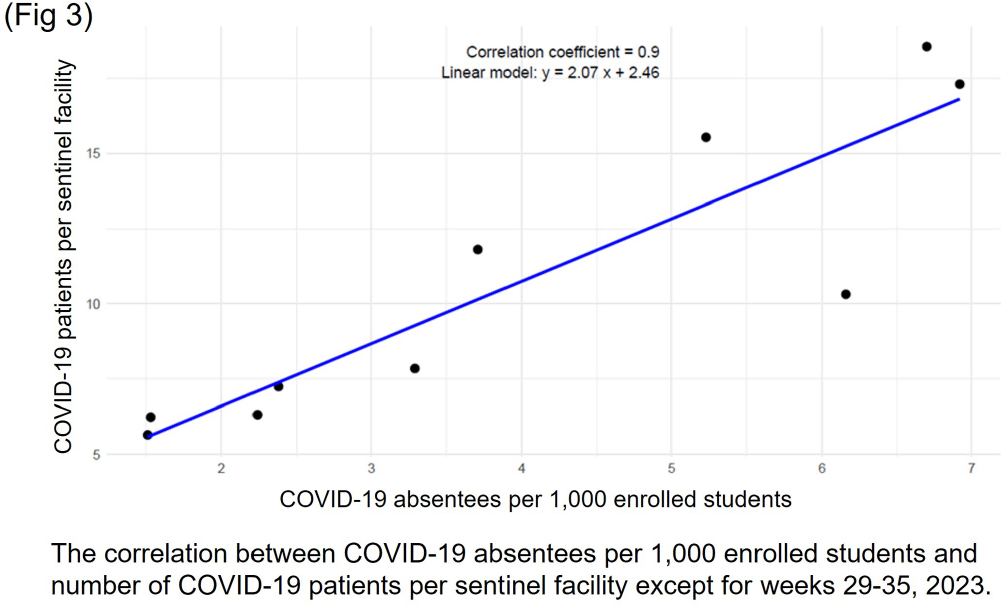
Background: In Japan, notifiable infectious disease surveillance ended and was replaced by sentinel surveillance following the COVID-19 reclassification in May 2023. Since COVID-19 sentinel surveillance is integrated into seasonal influenza surveillance, the number of reported cases varies depending on the extent to which sentinel facilities provide COVID-19 care. Therefore, we compared COVID-19 sentinel surveillance with school absentee surveillance, which is limited to high school equivalent age or younger, but provides reliable information on absences in the target population. Method: The 17-week period from week 23 (June 5 to June 11) to week 39 (September 25 to October 1) of 2023 was used as the target period. The number of weekly COVID-19 reports from 72 sentinel sites in Mie Prefecture (Population 1.7 million) as for sentinel surveillance and the number of COVID-19 absentees at a total of 998 facilities (401 kindergartens and nursery schools, and 597 elementary, junior high, and senior high schools) registered for school absentee surveillance in Mie Prefecture as for school absentee surveillance were compared across Mie Prefecture and eight health centers (Fig 1). Result: Except for the summer vacation period from week 29 to 35, sentinel surveillance and school absentee surveillance showed a significant positive correlation. During the summer vacation period, a decrease in the number of COVID-19 absentees was observed, especially in the elementary, junior high, and senior high school groups of the school absentee surveillance, compared to the sentinel surveillance (Fig 2 and 3). When compared by health center, no regional differences were observed in school absentee surveillance, but in sentinel surveillance, some health centers reported significantly more cases than others. Conclusion: The results of this study suggest that although COVID-19-based school absentee surveillance has some drawbacks, such as the limited number of subjects and the difficulty of evaluation during the summer vacation when schools are closed, it has the advantage of being able to evaluate the entire community without being affected by medical institution practice bias, and can be used to monitor trends in infectious diseases. It was considered important to combine and evaluate multiple surveillance indicators in order to accurately monitor epidemiologic trends of infectious disease over time.
Ensuring Accuracy: Making the Case for Inter-rater Reliability in Hospital Acquired Infection Surveillance
- Kelly Holmes, Mishga Moinuddin, Sandi Steinfeld
-
- Published online by Cambridge University Press:
- 16 September 2024, pp. s147-s148
-
- Article
-
- You have access Access
- Open access
- Export citation
-
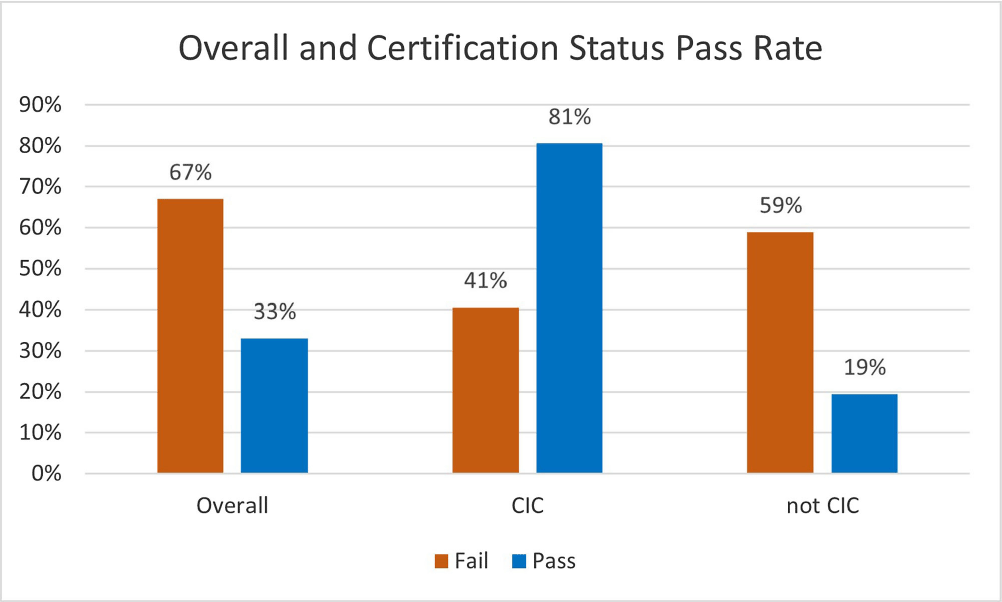
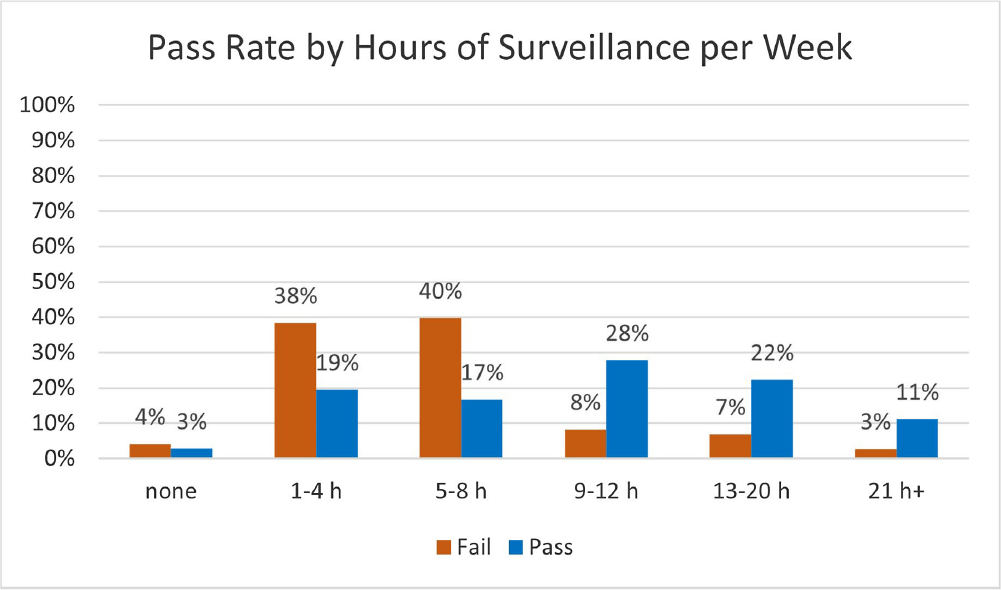
Background: Infection preventionists (Ips) have self-reported surveillance as the most time-consuming job task (1,2). APIC’s MegaSurvey 2020 reported that 60% of Ips consider themselves proficient or expert within this competency domain (2). Accurate coding of health care acquired infections is critical to identifying epidemiologically significant events, using data to improve practice, and compliance with state and federal mandated CMS reporting (3,4). Validated case study scenarios were distributed to infection preventionists to better understand how experience level and time spent performing surveillance affects interrater reliability (IRR) in applying the National Healthcare Safety Network (NHSN) surveillance definitions (5,6). Methods: Case study scenarios determined to have high item discrimination were added to an online test bank and distributed annually to Ips of varying experience levels and care settings as part of a mandatory training program (5). The test bank currently consists of forty-two validated questions. Each year, the participants receive approximately thirty questions, including twenty randomly selected from the test bank and ten beta scenarios under development. Only validated test bank scenarios are used to calculate the passing score of 85%. Participants are blinded to which questions are test bank scenarios versus beta scenarios. Additional information was gathered at the beginning of the test to determine CIC status, years of experience, and weekly hours spent doing surveillance. Data was analyzed for passing score on the first attempt for testing years 2019, 2021, 2022, and 2023. Results: Thirty-six Ips passed the IRR test on the first attempt (33%). Of those who passed on the first attempt, twenty-nine (81%) were certified and twenty-two (61%) reported at least nine hours a week performing surveillance. Of the seventy-three Ips (67%) who did not pass on the first attempt, thirty were certified (41%) and sixty (82%) reported performing surveillance for 8 hours or less per week. Conclusion: The first-time pass rate for certified and non-certified Ips was 33%, markedly lower than the self-reported proficiency rate of 60%. The majority of Ips who passed on the first attempt were certified and spent at least nine hours per week performing surveillance. certified and non-certified Ips who did not regularly perform surveillance as part of their weekly job tasks were less likely to pass the test on the first attempt. Given the first-time pass rate among all participants is below optimal, establishing inter-rater reliability systems and ongoing surveillance education for Ips is crucial to ensure accuracy of publicly reported data.
Community-associated Carbapenem-Resistant Organism Case Investigations in New York City, December 2020-May 2023
- Celina Santiago, Ying Lin, Ulrike Siemetzki-Kapoor, Nicole Burton, Katelynn Devinney, Balan Dominique, Portier Thomas, William Greendyke, Molly Kratz, Catharine Prussing, Kailee Cummings, Rebecca Zimba, Karen Alroy
-
- Published online by Cambridge University Press:
- 16 September 2024, p. s147
-
- Article
-
- You have access Access
- Open access
- Export citation
-
Epidemiology of carbapenem-resistant organisms (CRO) has focused on transmission in acute care hospital or long-term care facility (LTCF) settings. Few investigations have examined community-associated (CA)-CRO, with no consensus about common exposures. To explore possible exposures, the New York City (NYC) Department of Health and Mental Hygiene investigated suspected CA-CRO cases through routine surveillance among NYC residents with specimens collected during December 2020-May 2023. CA-CRO cases were defined as urine or skin specimens with bacterial cultures exhibiting carbapenem resistance, among individuals aged ≤70 years with no international travel, hospitalization, or LTCF stays within 12 months before specimen collection. Inclusion was determined by reviewing data from health information exchanges, when available electronic medical records, and telephone screening for those not excluded through record review. We identified 426 suspected cases for review, those not meeting the case definition were excluded; 44 individuals were not reached for screening. A preliminary questionnaire was fielded with 12 individuals and then refined to capture additional potential exposures. Analyses were completed with 23 individuals interviewed with the refined questionnaire. Of the 23, 70% were female; 39% were Hispanic, 17% Black, and 17% White; their median age was 60 years (range: 26-70 years). Further, 83% reported an outpatient appointment, 48% reported an outpatient procedure/surgery, and 9% reported having a hospitalized household member, all within 12 months before specimen collection; 26% had a urinary catheter or indwelling device within 2 days of specimen collection. Additionally, 30% reported taking antibiotics within 3 months of specimen collection, 52% denied taking antibiotics, 9% were unsure about antibiotic use, and 9% did not answer the question. Whole genome sequencing (WGS) was performed on 14 available isolates from CA-CRO cases by the NYC Public Health Laboratory or Wadsworth Center (WC), of which only 7 could be compared with isolates previously sequenced at WC (2017-2023). Six isolates were separated by >50 mutation events, suggesting no close genomic relationship. One isolate from 2021 was 11 mutation events from a 2018 isolate from the same individual, consistent with the expected evolutionary rate. While infrequent, CA-CRO cases occur in NYC. Outpatient healthcare, antibiotic use, and urinary catheters or indwelling devices were common self-reported exposures. Analyses were limited by screening non-response. Increased specimen availability for WGS could enhance investigation of CA-CRO exposure patterns. Health information exchange data were often incomplete and future surveillance could benefit from healthcare and public health partnerships and better documentation for more complete electronic medical histories.
Candida auris Screening of High-Risk Patients: A Descriptive Comparison of 2 Strategies.
- Laura Pedersen, Aldo Barajas-Ochoa, Kaila Cooper, Jenna Price, Kathryn Hannum, Yvette Major, Patrick R Ching, Barry Rittmann, Michelle Doll
-
- Published online by Cambridge University Press:
- 16 September 2024, p. s147
-
- Article
-
- You have access Access
- Open access
- Export citation
-
Background: Candida auris infection is associated with high morbidity and mortality. C. auris can persist in the healthcare environment and is associated with outbreaks. We compare screening strategies for C. auris in two high-risk patient populations. Methods: Our center is a tertiary, 865-bed hospital. In the context of known regional outbreaks of C. auris in post-acute care (PAC) facilities, we experienced extended clusters of apparent C. auris acquisition across several hospital units. Hospital acquisition was defined as new C. auris in clinical cultures in patients with no known history of C. auris colonization/infection. We performed point prevalence surveys (PPS) on affected units weekly until all tests were negative for two consecutive weeks. We also initiated admission screening for C. auris for patients admitted from PAC. All screening swabs were collected per CDC’s procedure. Tests were performed either by RT-PCR or Chromagar C. auris media, depending on availability. We compared the overall positivity rates of exposure PPS versus PAC admission screenings using Z-test for two proportions with statistical significance set at p < 0 .05 Results: From 2/2023-12/2023, a total of 533 tests on 367 unique patients were processed during PPS; 512 tests were negative and 21 were positive (3.9% positivity rate). Three additional samples were either unable to be processed or indeterminate. There were 68 patients who had repeat testing weekly for ≥2 weeks. Most remained negative, but 5 tested positive after variable amounts of negative-week intervals: 3 patients at week 2, 1 patient at week 4 and 1 patient at week 5. From 8/2023 to 12/2023, a total of 89 patients admitted from 35 different PAC facilities underwent admission screening for C. auris. Only three patients were positive (3.4%), each from a different facility. The difference in the positivity rates between PPS and PAC was not statistically significant (Z-score 0.25, p = 0.79). Discussion: Our C. auris screening strategies found similar positivity rates for patients admitted to the hospital from PACs compared to targeted PPS in the setting of apparent hospital acquisition events. These strategies may be considered as complementary. Facilities experiencing apparent acquisition events should consider screening high-risk admissions to identify and isolate colonized patients, particularly if standard infection prevention practices are being performed with high fidelity.






















