Introduction
Milk and milk products are important global dietary products consumed by more than 6 billion people worldwide. In 2021, recorded milk consumption was 928 million tons, making the dairy industry a very profitable market (Food and Agriculture Organization of the United Nations, 2021). However, an infection of the mammary gland caused mainly by bacteria, mastitis, is a major problem affecting animal welfare, productivity and the economy, especially in dairy production, which can lead to losses for the dairy industry (Abera et al., Reference Abera, Demie, Aragaw, Regassa and Regassa2013).
Among the main microorganisms involved in mastitis, we can mention Staphylococcus sp., of which S. aureus is the most significant mastitis pathogen, whilst the NAS (non-aureus staphylococci) are regarded as minor mastitis pathogens. Among all NAS found among cattle, S. haemolyticus, S. chromogenes, S. epidermidis, S. warneri, S. cohnii, S. simulans, S. hominis, S. capitis, and S. xylosus are the most prevalent species (Traversari et al., Reference Traversari, Van Den Borne, Dolder, Thomann, Perreten and Bodmer2019). Besides NAS, it is important to highlight the new genus reclassified from Staphylococcus, the Mammaliicoccus. A study published by Madhaiyan et al. (Reference Madhaiyan, Wirth and Saravanan2020) suggests the reassignment of five species of Staphylococcus (S. sciuri, S. fleurettii, S.lentus, S. stepanovicii and S. vitulinus) to this novel genus, with Mammaliicoccus sciuri as the type species. By whole genome sequences, these mammaliicocci species are as distant from staphylococci (64.2–67.3% amino acid identity – AAI) as Macroccus is distant from Staphylococcus (61.1–64.3%). Non-aureus staphylococci and mammaliicocci (NASM) have become a concern among dairy producers, as they are the most common microorganisms isolated from aseptically collected quarter milk samples (Fergestad et al., Reference Fergestad, Touzain, De Vliegher, De Visscher, Thiry, Ngassam Tchamba, Mainil, Abee-Lund, Blanchard and Wasteson2021).
According to the World Health Organization (WHO, 2024) 420 000 lives are lost due to food poisoning, and Staphylococcus spp. are characterized as important agents that can cause foodborne diseases. Poisoning occurs due to ingesting enterotoxins produced in food, including those produced by NASM, and symptoms include vomiting, diarrhea and cramps (da Silva Cândido et al., Reference da Silva Cândido, da Silva, de Matos, da Silva do Nascimento, Camargo, Cobo Zanella, Mores Rall and Silva2020). But there are other problems as well. Political agendas, legislation, development of therapies and educational initiatives are essential to mitigate the increasing rate of antibiotic resistance amongst pathogenic organisms. In addition to bacteria having the ability to form biofilms, other genes are also involved in antibiotic resistance, and the process of becoming resistant can also occur through mutation at the genetic level (Zaman et al., Reference Zaman, Bin Hussain, Nye, Mehta, Mamun and Hossain2017). In addition, it has been suggested that NASM might act as a potential reservoir for resistance genes, which can be transferred and integrated into the genome of S. aureus (Vitali et al., Reference Vitali, Petrelli, Lamikanra, Prenna and Akinkunmi2014). Despite being under debate, it is essential to highlight that there are species of Mammaliicoccus that are suggested to be the evolutionary origin of the mecA gene in S. aureus (Lakhundi and Zhang, Reference Lakhundi and Zhang2018).
For the most part, NASM can be regarded as minor pathogens that do have the ability to cause mastitis. However, some studies have demonstrated that NAS can reduce infection by S. aureus and other pathogens. Therefore, this review article aims to address the negative and positive aspects that NASM can have in bovine mastitis, in addition to addressing the importance of adopting the concept of One Health.
Non-aureus staphylococci
Differences among and within bovine NAS species have been reported regarding epidemiology and ecology (Souza et al., Reference Souza, Piepers, Della Libera, Heinemann, Cerqueira and De Vliegher2016), antimicrobial resistance (Fergestad et al., Reference Fergestad, Touzain, De Vliegher, De Visscher, Thiry, Ngassam Tchamba, Mainil, Abee-Lund, Blanchard and Wasteson2021), virulence (Wuytack et al., Reference Wuytack, De Visscher, Piepers, Boyen, Haesebrouck and De Vliegher2020), potential protective traits (Toledo-Silva et al., Reference Toledo-Silva, de Souza, Mertens, Piepers, Haesebrouck and De Vliegher2021a, Reference Toledo-Silva, Nogueira De Souza, Piepers, Mertens, Haesebrouck and De Vliegher2021b), host interaction (Piccart et al., Reference Piccart, Verbeke, De Visscher, Piepers, Haesebrouck and De Vliegher2016) and impact on udder health and milk yield (Valckenier et al., Reference Valckenier, Piepers, De Visscher, Bruckmaier and De Vliegher2019, Reference Valckenier, Piepers, De Visscher and De Vliegher2020, Reference Valckenier, Piepers, Schukken, De Visscher, Boyen, Haesebrouck and De Vliegher2021) as recently reviewed by De Buck et al. (Reference De Buck, Ha, Naushad, Nobrega, Luby, Middleton, De Vliegher and Barkema2021). Non-aureus staphylococci can be present in a broad range of ecological habitats, which includes bovine teat canals (Traversari et al., Reference Traversari, Van Den Borne, Dolder, Thomann, Perreten and Bodmer2019) and teat apices (Adkins et al., Reference Adkins, Dufour, Spain, Calcutt, Reilly, Stewart and Middleton2018) as well as the farm environment including bulk tank milk, bedding (Adkins et al., Reference Adkins, Placheta, Borchers, Bewley and Middleton2022) and feces (Wuytack et al., Reference Wuytack, De Visscher, Piepers, Boyen, Haesebrouck and De Vliegher2020). S. chromogenes and S. epidermidis are predominantly isolated from milk. Therefore, they are considered host-adapted species, whereas S. equorum and S. devriesei are mostly found in the environment and are classified as environmental NAS species (Adkins et al., Reference Adkins, Placheta, Borchers, Bewley and Middleton2022). Other species, such as S. haemolyticus, originate from various (extra)mammary sites and so are labeled opportunistic (Adkins et al., Reference Adkins, Placheta, Borchers, Bewley and Middleton2022). Unfortunately, multiple studies contradict this current species-stratification (Thorberg et al., Reference Thorberg, Danielsson-Tham, Emanuelson and Persson Waller2009; Supré et al., Reference Supré, Haesebrouck, Zadoks, Vaneechoutte, Piepers and De Vliegher2011; De Visscher et al., Reference De Visscher, Supré, Haesebrouck, Zadoks, Piessens, Van Coillie, Piepers and De Vliegher2014; Vanderhaeghen et al., Reference Vanderhaeghen, Piepers, Leroy, Van Coillie, Haesebrouck and De Vliegher2015), and these confounding observations could be attributed to strain differences within NAS species. Wuytack et al. (Reference Wuytack, De Visscher, Piepers, Boyen, Haesebrouck and De Vliegher2019), for example, suggested in a recent study on strain distribution of NAS in multiple bovine-associated habitats that different NAS strains within a species appear to be habitat-specific or site-specific.
After several taxonomic reclassifications, introducing the coagulase test enabled differentiation between CoNS and CoPS (Fairbrother, Reference Fairbrother1940). Non-aureus staphylococci is a group of Gram-positive bacteria that were before known as coagulase-negative staphylococci (CoNS); they are usually found in the normal flora of humans and different animals, and some species are recognized as human and facultative animal pathogens (Belhout et al., Reference Belhout, Elgroud and Butaye2022). For a long time, only S. aureus was considered to be harmful to its host, however, as a consequence of patient- and procedure-related changes, NAS now includes a large number of nosocomial microorganisms, with S. epidermidis and S. haemolyticus being the most significant ones (Pereira-Ribeiro et al., Reference Pereira-Ribeiro, Sued-Karam, Faria, Nogueira, Colodette, Fracalanzza, Duarte, Júnior and Mattos-Guaraldi2019).
Some of the pathogenicity of NAS strains is probably due to their ability to adhere to surfaces and form biofilm on polymer surfaces of medical devices used during the hospitalization process in hospitals (Le et al., Reference Le, Villaruz, Zheng, He, Fisher, Nguyen, Ho, Yeh, Joo, Cheung and Otto2019). In addition, NAS were also recognized to be capable of ‘hiding’ from the immune system by being internalized by nonprofessional phagocytes, a mechanism that is mediated by the AtlE gene that helps the microorganism not only bind but also be phagocyted by human endothelial cells (Christner et al., Reference Christner, Franke, Schommer, Wendt, Wegert, Pehle, Kroll, Schulze, Buck, MacK, Aepfelbacher and Rohde2010). Besides that, NAS showed antimicrobial resistance and carried multiple resistance genes, which can be shared with other species (Lee et al., Reference Lee, Kim, Do Park and Yang2020).
NAS brings issues not only to human health but also to animals. They are often found as the majority in bovine intramammary infection, especially in dairy heifers, and several studies in different areas of the globe have found NAS in different proportions in bovine udder, mainly in non-bred or first lactation animals (De Buck et al., Reference De Buck, Ha, Naushad, Nobrega, Luby, Middleton, De Vliegher and Barkema2021). This could have happened because the NAS group was considered to comprise minor pathogens and so were typically not treated. However, grouping these species contributed to a better understanding of their distribution and importance to the animal's health care (Condas et al., Reference Condas, De Buck, Nobrega, Carson, Roy, Keefe, DeVries, Middleton, Dufour and Barkema2017).
Despite NAS being a large group, we can highlight some individual species. S. epidermidis is recognized as a pathogen and is the most important cause of nosocomial infection (Hugh and Ellis, Reference Hugh and Ellis1968). This species can also adapt quickly to extreme conditions. For example, to survive in a place with a high salt concentration, the species has a different number of ion/proton exchangers and systems for osmo protectants (Gill et al., Reference Gill, Fouts, Archer, Mongodin, DeBoy, Ravel, Paulsen, Kolonay, Brinkac, Beanan, Dodson, Daugherty, Madupu, Angiuoli, Durkin, Haft, Vamathevan, Khouri, Utterback and Fraser2005), and carries many genes that allow it to survive in these severe situations (Otto, Reference Otto2009). Also, it is known that S. epidermidis is capable of producing an extracellular polysaccharide adhesin, termed polysaccharide intercellular adhesin (PIA) or polymeric N-acetyl-glucosamine, by ica operon-encoded enzymes. This is a significant virulence factor of the specie, helping them to colonize medical devices and leading to device-related infections (Le et al., Reference Le, Villaruz, Zheng, He, Fisher, Nguyen, Ho, Yeh, Joo, Cheung and Otto2019). Besides this, they can also produce different toxins (Namvar et al., Reference Namvar, Bastarahang, Abbasi, Ghehi, Farhadbakhtiarian, Arezi, Hosseini, Baravati, Jokar and Chermahin2014). However, studies suggest that this bacterium has evolved to play an important role in maintaining the microbiomes of humans and other mammals (Otto, Reference Otto2009).
Due to its potential to produce biofilm, S. haemolyticus is also a critical species for nosocomial infections in humans, especially from blood infections. However, there is not as much information for them as for S. epidermidis (Barros et al., Reference Barros, Lemos, Souto-Padrón and Giambiagi-deMarval2015). When the resistance to β-lactam antibiotics is considered, S. aureus and S. haemolyticus present enough similarity to say that they have the exact same resistance mechanism (Czekaj et al., Reference Czekaj, Ciszewski and Szewczyk2015).
In several countries, S. chromogenes has been identified as the most prevalent species in milk samples from healthy cows and those with subclinical mastitis and clinical mastitis (Valckenier et al., Reference Valckenier, Piepers, De Visscher and De Vliegher2020). S. chromogenes could overcome the physical barrier of the udder and form biofilms. In addition, this species is more adapted to the cow's mammary gland and may become a microbial reservoir and possible source of infection (Tomazi et al., Reference Tomazi, Gonçalves, Barreiro, Arcari and dos Santos2015; Isaac et al., Reference Isaac, Bohl, Breser, Orellano, Conesa, Ferrero and Porporatto2017). Molecular studies have also shown that S. chromogenes has several antibiotic resistance genes, in addition to having this resistance tested and confirmed in phenotypic tests (Qu et al., Reference Qu, Zhao, Nobrega, Cobo, Han, Zhao, Li, Li, Barkema and Gao2019).
Mammaliicoccus
Mammaliicoccus are Gram-positive, non-motile, non-spore-forming, paired cocci cells with aerobic to facultative anaerobic growth. They are catalase-positive and possess variable oxidase activities. The G + C content varies from 31.6 to 35.7% in genomes that range from 2.44 to 2.81 Mbp. Based on the analyses of 16S rRNA gene sequences, five species (M. sciuri, M. fleurettii, M. lentus, M. stepanovicii, and M. vitulinus), are now in this novel genus (Madhaiyan et al., Reference Madhaiyan, Wirth and Saravanan2020) and are as distant from Staphyloccus species as Macrococcus (another genus) is from Staphylococcus. Recognized to colonize humans and animals, they are also considered potentially opportunist pathogens capable of affecting the immune system (Beuckelaere et al., Reference Beuckelaere, De Visscher, Souza, Meyer, Haesebrouck, Piepers and De Vliegher2021). Like the NAS, this group may also cause mastitis in different species, and/or may serve to transmit resistance genes to other more pathogenic species, as suggested by the study of Lienen et al. (Reference Lienen, Schnitt, Hammerl, Maurischat and Tenhagen2022) where M. sciuri, M. lentus, M. fleurettii and M. vitulinus were isolated from milk and swab samples collected from German dairy cows.
M. sciuri (previously Staphylococcus sciuri) is usually found on the skin of wild and domestic animals. Based on their ribotype pattern, the species was divided into three different subspecies called S. sciuri subsp. sciuri, S. sciuri subsp. carnaticus and S. sciuri subsp. rodentium (Nemeghaire et al., Reference Nemeghaire, Argudín, Feßler, Hauschild, Schwarz and Butaye2014). In addition, some studies suggest this species as the origin of the mecA gene, which is located on a mobile genetic element called staphylococcal cassette chromosome mec (SCCmec), and is the gene responsible for methicillin-resistance in S. aureus (MRSA) or more generally in Staphylococcus spp (MRSS: Couto et al., Reference Couto, De Lencastre, Severina, Kloos, Webster, Hubner, Sanches and Tomasz1996; Fuda et al., Reference Fuda, Suvorov, Shi, Hesek, Lee and Mobashery2007). Whilst being characterized as a typical colonizer of a wide range of animals, this species has also been associated with various health problems including skin disease in goats and pigs (Nemeghaire et al., Reference Nemeghaire, Argudín, Feßler, Hauschild, Schwarz and Butaye2014). Despite displaying various virulence factors (lipolytic, proteolytic and hemolytic), these bacteria are not related to infection in humans, except after previous infection due to a pathogen (Nemeghaire et al., Reference Nemeghaire, Argudín, Feßler, Hauschild, Schwarz and Butaye2014).
M. lentus (previously S. lentus) was first characterized as a subspecies of S. sciuri. However, after analysis of DNA-DNA hybridization, it was placed into a new species (Schleifer et al., Reference Schleifer, Geyer, Kilpper-Bälz and Devriese1983). It was originally isolated from poultry and other animals and can produce enterotoxins, but only to a minor extent (less than 100 ng/ml supernatant: Irlinger, Reference Irlinger2008). Guerre et al. (Reference Guerre, Bailly, Benard and Burgat1999) showed that the enterotoxigen strain of M. lentus inoculated in milk for the production of Camembert-type cheeses does not produce enterotoxins despite high levels of the microorganisms.
For M. fleurettii (previously S. fleurettii), it was also possible to identify enterotoxin production, and some studies suggest that this species, rather than S. sciuri, is the origin of the mecA gene in MRSA. However, the exact origin is still debated (Lakhundi and Zhang, Reference Lakhundi and Zhang2018). In this context, M. stepanovicii was also reported with the presence of the gene (Loncaric et al., Reference Loncaric, Kübber-Heiss, Posautz, Stalder, Hoffmann, Rosengarten and Walzer2013). Table 1 provides a complete list of NASM and the different samples from which they were isolated.
Table 1. NASM isolated from different samples (human, animal, environment, and food) and different countries

NASM as a cause of mastitis
NAS forms a heterogeneous group of more than 50 species, of which approximately ten are associated with intramammary infections in dairy cattle and are abundantly present in bulk milk (Vanderhaeghen et al., Reference Vanderhaeghen, Piepers, Leroy, Van Coillie, Haesebrouck and De Vliegher2015). As other species of Staphylococci become more commonly associated with the disease, it is of interest to learn more about the distribution of NAS on dairy farms. They have many virulence factors, and mastitis control is complicated because this group contains many species (Thorberg et al., Reference Thorberg, Danielsson-Tham, Emanuelson and Persson Waller2009).
The main ways to diagnose subclinical mastitis from NAS are from the results of somatic cell count (SCC) and culture. Tomazi et al. (Reference Tomazi, Gonçalves, Barreiro, Arcari and dos Santos2015) observed that mammary quarters infected by NAS had elevated SCC (> 306 000 vs. < 65 000 in contralateral uninfected quarters), without any affect on milk yield or contents of fat, crude protein, casein, lactose, total solids, and solids-not-fat. Although considered as pathogens with a lower clinical incidence, NAS infections have been identified as relatively common (Persson Waller et al., Reference Persson Waller, Aspán, Nyman, Persson and Grönlund Andersson2011; Gao et al., Reference Gao, Barkema, Zhang, Liu, Deng, Cai, Shan, Zhang, Zou, Kastelic and Han2017; Heikkilä et al., Reference Heikkilä, Liski, Pyörälä and Taponen2018), clinical cases being due mainly to S. chromogenes, S. simulans and S. haemolyticus (Persson Waller et al., Reference Persson Waller, Aspán, Nyman, Persson and Grönlund Andersson2011). In subclinical mastitis cases, NAS are the second most commonly identified pathogen after S. aureus in Swedish dairy cows, S. epidermidis followed by S. chromogenes, S. simulans, and S. haemolyticus being the most common (Persson Waller et al., Reference Persson Waller, Aspán, Nyman, Persson and Grönlund Andersson2011). Another study isolated S. chromogenes, S. fleurettii, S. haemolyticus, S. sciuri, S. simulans, S. succinus and S. xylosus from mastitic bovine milk (Khazandi et al., Reference Khazandi, Al-Farha, Coombs, O'Dea, Pang, Trott, Aviles, Hemmatzadeh, Venter, Ogunniyi, Hoare, Abraham and Petrovski2018).
The main pathogenic factors of NASM
According to Vanderhaeghen et al. (Reference Vanderhaeghen, Piepers, Leroy, Van Coillie, Haesebrouck and De Vliegher2014), there are three virulence factor groups involved in the pathogenesis of staphylococcal infections: secreted proteins (e.g., superantigens, cytotoxins and tissue-degrading enzymes), cell surface-bound proteins (e.g. microbial surface components recognizing adhesive matrix molecules: MSCRAMM) and cell wall components, with the polysaccharide capsule and lipoteichoic being examples of those. Naushad et al. (Reference Naushad, Naqvi, Nobrega, Luby, Kastelic, Barkema and De Buck2019), using whole genome sequencing analyzed the distribution of 191 virulence factors in 441 genomes of 25 NAS species; the authors reported virulence factors in functional categories such as toxins (n = 93), iron metabolism (n = 29), adherence (n = 28), exoenzymes (n = 21) and immune evasion (n = 20). Another study by Wuytack et al. (Reference Wuytack, De Visscher, Piepers, Boyen, Haesebrouck and De Vliegher2020) compared the presence of virulence genes (agrA, bap, and cap5H and the mecA methicillin resistance gene) of NAS isolated from the milk samples of healthy quarters and quarters with clinical mastitis. Based on this comparison, they suggested that NAS has the potential to cause mild clinical mastitis.
Enterotoxins
Staphylococcal poisoning is widespread and mainly transmitted by food products, as it can affect healthy people. NAS is also able to produce enterotoxins, which can be one of the causes of many food diseases (da Silva Cândido et al., Reference da Silva Cândido, da Silva, de Matos, da Silva do Nascimento, Camargo, Cobo Zanella, Mores Rall and Silva2020). Enterotoxins belong to a group of extracellular single-chain proteins that are water-soluble and resistant to proteolytic enzyme action in the digestive system. This means that they remain active after ingestion and can compromise human health and, consequently, the public health system. These proteins are also thermostable and resist thermal treatments such as pasteurization (Vanderhaeghen et al., Reference Vanderhaeghen, Piepers, Leroy, Van Coillie, Haesebrouck and De Vliegher2014). There are more than 25 types of staphylococcal enterotoxins (SE) and enterotoxin-like (SEL) toxins; the former have an emetic effect in primates, whilst the latter do not. The most known of the enterotoxins are the classical serotypes (SEA through SEE: Bergdoll et al., Reference Bergdoll, Borja, Robbins and Weiss1971), followed by the non-classical SEG through SET as well as SEY and more recently SE02 (Su and Lee Wong, Reference Su and Lee Wong1995; Munson et al., Reference Munson, Tremaine, Betley and Welch1998; Ono et al., Reference Ono, Omoe, Imanishi, Iwakabe, Hu, Kato, Saito, Nakane, Uchiyama and Shinagawa2008, Reference Ono, Sato'o, Narita, Naito, Hirose, Hisatsune, Asano, Hu, Omoe, Sugai and Nakane2015, Reference Ono, Hirose, Narita, Sugiyama, Asano, Hu and Nakane2019; Omoe et al., Reference Omoe, Hu, Ono, Shimizu, Takahashi-Omoe, Nakane, Uchiyama, Shinagawa and Imanishi2013; Suzuki et al., Reference Suzuki, Ono, Shimojima, Kubota, Kato, Kakuda, Hirose, Hu, Nakane, Takai and Sadamasu2020). The SELs identified until today are SELJ and then SELU through SELX, SELZ and SEL 01 (Zhang et al., Reference Zhang, Iandolo and Stewart1998; Letertre et al., Reference Letertre, Perelle, Dilasser and Fach2003; Thomas et al., Reference Thomas, Jarraud, Lemercier, Cozon, Echasserieau, Etienne, Gougeon, Lina and Vandenesch2006; Wilson et al., Reference Wilson, Seo, Cartwright, Connelley, Chuang-Smith, Merriman, Guinane, Park, Bohach, Schlievert, Morrison and Fitzgerald2011; Spoor et al., Reference Spoor, Richardson, Richards, Wilson, Mendonca, Gupta, McAdam, Nutbeam-Tuffs, Black, O'gara, Lee, Corander and Ross Fitzgerald2015; Hisatsune et al., Reference Hisatsune, Hagiya, Shiota and Sugai2017). NASM are capable of producing the majority of these (Rodrigues et al., Reference Rodrigues, Silva, Trevilin, Cruzado, Mui, Duarte, Castillo, Canniatti-Brazaca and Porto2017; da Silva Cândido et al., Reference da Silva Cândido, da Silva, de Matos, da Silva do Nascimento, Camargo, Cobo Zanella, Mores Rall and Silva2020). However, this has largely been ignored by authorities as NAS can establish a commensal relationship with humans and animals since they are also part of their normal microbiota (Podkowik et al., Reference Podkowik, Park, Seo, Bystroń and Bania2013).
When de Freitas Guimarães et al. (Reference de Freitas Guimarães, Nóbrega, Richini-Pereira, Marson, de Figueiredo Pantoja and Langoni2013) studied 263 isolates of Staphylococcus spp. from mastitic bovine milk samples they were able to show that 49% were NAS, and of these 66% contained the gene responsible for enterotoxin production, with sea, seb and sec (producing the equivalent classical SEs) being the ones that appeared most. In addition, Ünal and Çinar (Reference Ünal and Çinar2012) found that 46% of isolated NAS tested positive for having one or more SE genes, with 63% out of the total presenting the seh-sej gene, which belongs to the non-classical SEs. Corroborating these results, Rodrigues et al. (Reference Rodrigues, Silva, Trevilin, Cruzado, Mui, Duarte, Castillo, Canniatti-Brazaca and Porto2017) characterized NAS isolated from raw milk, Minas cheese, and Minas cheese production lines. They detected the genes seb, seg, seh, sek and seu in the NAS species identified.
Biofilm formation
Different species of bacteria can live in communities, adhere to an inert or living surface, and be protected by a polymeric matrix called a biofilm. This matrix is a self-product of its metabolism, consisting mainly of sugar and proteins (Shemesh and Ostrov, Reference Shemesh and Ostrov2020). Biofilm formation in staphylococcal species is associated with a number of factors, and several studies in the literature show NAS as carriers of biofilm genes. Production of PIA is mediated by the genes icaA, icaB, icaC and icaD, representing an important contribution to cell–cell adhesion (Srednik et al., Reference Srednik, Tremblay, Labrie, Archambault, Jacques, Cirelli and Gentilini2017). One study observed biofilm formation in 50% of the NAS strains studied (Rumi et al. (Reference Rumi, Huguet, Bentancor and Gentilini2013). Naushad et al. (Reference Naushad, Naqvi, Nobrega, Luby, Kastelic, Barkema and De Buck2019), in a study based on whole-genome sequencing data for 441 isolates from 25 species, identified that the icaC gene of the ica operon, believed to be involved in biofilm formation, was the second most frequent gene, and it was present in 17 isolates out of all species studied. Other genes of the ica operon, namely icaA, were found in eight species, icaB in seven species and icaD in eight.
Bap is another protein involved in biofilm formation, which is responsible for adhesion. In their research, Piessens et al. (Reference Piessens, De Vliegher, Verbist, Braem, Van Nuffel, De Vuyst, Heyndrickx and Van Coillie2012) showed that 11.2% (41/366) of NAS were bap gene positive. Zuniga et al. (Reference Zuniga, Melville, Saidenberg, Laes, Gonsales, Salaberry, Gregori, Brandão, dos Santos, Lincopan and Benites2015) isolated 67 strains of NAS from subclinical bovine mastitis and tested for the presence of the bap gene; they reported that 45 (67.2%) isolates tested positive and could thus produce biofilms. Rohde et al. (Reference Rohde, Burandt, Siemssen, Frommelt, Burdelski, Wurster, Scherpe, Davies, Harris, Horstkotte, Knobloch, Ragunath, Kaplan and Mack2007) found that nearly 90% of S. epidermidis strains produced a biofilm, and up to 27% of biofilm-positive S. epidermidis isolates produced PIA-independent biofilms, which involves the expression and accumulation of the associated protein. This indicates that, besides Bap, other proteins are involved in ica-idependent biofilm formation. Also, NAS produces MSCRAMM proteins, which mediate staphylococcal adherence to the host's extracellular matrix components. In addition to the operon ica that produces PIA adhesion, these components also form a biofilm (Christner et al., Reference Christner, Franke, Schommer, Wendt, Wegert, Pehle, Kroll, Schulze, Buck, MacK, Aepfelbacher and Rohde2010).
Microbial surface components recognizing adhesive matrix molecules (MSCRAMM)
Adhesion to host proteins is usually mediated by bacterial cell-wall-associated proteins called MSCRAMMs (Walsh et al., Reference Walsh, Miajlovic, Gorkun and Foster2008). A gene related to biofilm formation is embp, which is a MSCRAMM that plays a role during primary attachment to conditioned surfaces (Christner et al., Reference Christner, Franke, Schommer, Wendt, Wegert, Pehle, Kroll, Schulze, Buck, MacK, Aepfelbacher and Rohde2010). According to Turchi et al. (Reference Turchi, Bertelloni, Marzoli, Cerri, Tola, Azara, Longheu, Tassi, Schiavo, Cilia and Fratini2020), this was their study's most frequently detected gene, followed by eno and fbe, then altE. Srednik et al. (Reference Srednik, Tremblay, Labrie, Archambault, Jacques, Cirelli and Gentilini2017) showed in their research that NAS isolated from cows with mastitis can produce biofilm in vitro, and they also showed that 90% of the isolates tested positive for one of the above genes.
Zuniga et al. (Reference Zuniga, Melville, Saidenberg, Laes, Gonsales, Salaberry, Gregori, Brandão, dos Santos, Lincopan and Benites2015) also studied the components on microbial surfaces that recognize the adhesive matrix and found the presence of the can gene in 17% of isolates, and the eno gene at 82.1% was the most prevalent gene among all. The ebpS gene was positive in 17.9%, and there were 72.6% strains positive for the fnbA gene and 5.7% for the fnaB gene. The fib gene was among the most prevalent, present in 76/106 (71.6%), and the bap gene was in 60/106 (56%). Darwish and Asfour (Reference Darwish and Asfour2013) identified the presence of the eno gene in 92.6% of all 68 NAS isolates. Simojoki et al. (Reference Simojoki, Hyvönen, Plumed Ferrer, Taponen and Pyörälä2012) also showed a prevalence of the eno gene, which was positive in 75% of the 84 studied strains, and the genes bbp (1.2%), cflA (4.8%), cflB (3.6%), ebpS (4.8%), fib (4.8%), fnbA (3.6%), and fnbB (2.4%) were also detected. The cflA gene is involved with adhesion and immune evasion whilst the ebpS gene is responsible for encoding elastin-balding proteins present in the extracellular matrix. Fibrinogen-binding proteins are encoded by the fib gene, and fibronectin-binding protein A and fibronectin-binding protein B are encoded by the fnbA and fnbB genes respectively (Zuniga et al., Reference Zuniga, Melville, Saidenberg, Laes, Gonsales, Salaberry, Gregori, Brandão, dos Santos, Lincopan and Benites2015).
Resistance to antibiotics
Biofilm formation is also connected to antibiotic resistance. According to Srednik et al. (Reference Srednik, Tremblay, Labrie, Archambault, Jacques, Cirelli and Gentilini2017), NAS isolates growing within biofilms are less susceptible to antimicrobials commonly used on farms. Antibiotics are manufactured at an estimated scale of about 100 000 tons annually worldwide, and their use profoundly impacts bacterial life on Earth. More pathogenic strains have become antibiotic-resistant, and some have become resistant to multiple antibiotics, known as multidrug resistance (Nikaido, Reference Nikaido2009). El-Seedy et al. (Reference El-Seedy, Radwan, Hassan and Shehata2017) tested how biofilm formation could affect antibiotic resistance, and a total of 95 NAS were identified and tested for biofilm formation, of which 95.8% were resistant to ampicillin, 77.9% were resistant to cefoxitin, 35.8% were resistant to cefuroxime and 31.6% were resistant to amoxycillin.
Besides biofilm formation, there are also genes involved in antibiotic resistance. Xu et al. (Reference Xu, Tan, Zhang, Xia and Sun2015) studied the presence of some of these genes in 76 NAS isolates. They reported the following results: the most prevalent gene was linA, which gives resistance to lincosamides, followed by tetK, which is responsible for resistance to tetracycline. Conferring resistance to penicillin, the gene blaZ was found in 30.3% Also present were the genes aacA – aphD which provide resistance to aminoglycoside. Another gene studied was msrA/B, which was positive in 17.1 and 19.7% of isolates, respectively, conferring resistance to macrolide and erythromycin. The gene mecA is one of the most studied genes, and it is responsible for methicillin resistance and was positive in 17.1% of isolates; the genes ermB/C that confer resistance to erythromycin and clindamycin were present in 9.2 and 13.2% respectively whilst the genes aac (6′)/aph (2′), responsible for resistance to streptomycin, were present in 10.5% of the strains (Kumar et al., Reference Kumar, Yadav and Singh2010).
Another study identified 109 NAS, all of which were positive for the blaZ gene, and 73% of isolates had the mecA gene, both of which resist β-lactam groups. The genes that grant resistance to tetracycline tetK, tetL and tetM all showed a high presence, and other genes identified in a significant amount of isolates were the MLSb resistance genes, ermB, inuA, msrA, and mphC. In addition, dfrG, aacA-aphD and aphA3 were present in 25, 24, and 43 NAS isolates (Qu et al., Reference Qu, Zhao, Nobrega, Cobo, Han, Zhao, Li, Li, Barkema and Gao2019). Ahmed et al. (Reference Ahmed, Neubauer, Tomaso, El Hofy, Monecke, Abdeltawab and Hotzel2020) found NASM isolates exhibited a high prevalence of the resistance genes mecA, blaZ and tetK. Approximately half of the isolates harbored the aac-aphD, erm(C), and erm(B) genes.
Klibi et al. (Reference Klibi, Maaroufi, Torres and Jouini2018) studied 83 isolates; sixty-eight of these strains were identified as NAS. In the study, 29.41% showed oxacillin and cefoxitin resistance whislt some strains were mecA positive. Eleven of those twenty methicillin resistant non-aureus staphylococci (MRNAS) showed erythromycin-resistance and were positive for ermB, ermT, ermC, mphC or msrA. Four strains were tetK positive, which were resistant to tetracycline. Silva et al. (Reference Silva, Guimarães, Manzi, Júnior, Gómez-Sanz, Gómez, Langoni, Rall and Torres2014) also studied MRNAS, and they reported 26 strains with oxacillin and cefoxitin resistance that were positive for the mecA gene. In the same study, the SCCmec gene, a mobile genetic element that carries the central determinant for broad-spectrum beta-lactam resistance encoded by the mecA gene, was analyzed; it was present in 9 out of 26 strains. MRNAS isolates were also found to display additional resistance to tetracycline, streptomycin, tobramycin and gentamicin, and harbored the genes: tet(K), str, ant(4′)-I, and aac(6′)aph(2″).
Multidrug resistance
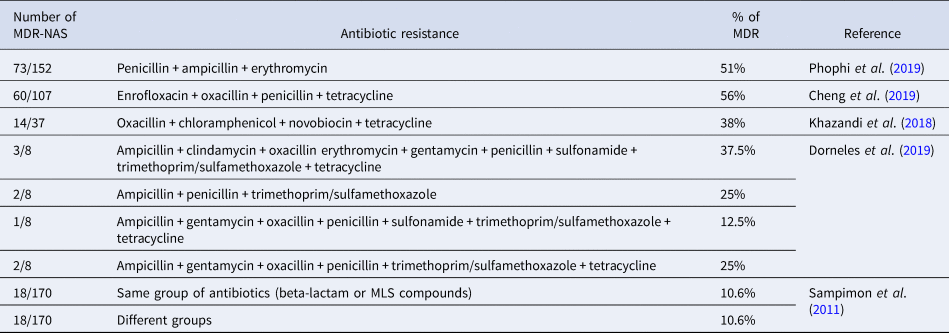
Studies suggest that the frequent use of drugs in veterinary practices, such as penicillin and tetracycline, especially among cattle, has increased the resistance of microorganisms over the years. The Food and Drug Administration (FDA) and the European Medicine Agency (EMA) show that tetracycline, penicillin, and sulfonamides are the classes of antibiotics most sold in the USA and Europe (Dorneles et al., Reference Dorneles, Fonseca, Abreu, Lage, Brito, Pereira, Brandão, Guimarães and Heinemann2019). Several studies in the literature point to multidrug-resistant NAS and these are enumerated in Table 2. In addition, NAS are known to be more resistant to antibiotics when compared to S. aureus, and this resistance can be to different drugs and contribute to a low cure rate. The resistance can be due to NAS, typically on the cow's body, being more exposed to antimicrobials (Khazandi et al., Reference Khazandi, Al-Farha, Coombs, O'Dea, Pang, Trott, Aviles, Hemmatzadeh, Venter, Ogunniyi, Hoare, Abraham and Petrovski2018).
Table 2. Number of multidrug resistant non-aureus Staphylococci; antibiotic groups they have resistance to; percentage of resistant strains for each drug or group.
NASM involvement in mastitis prevention
While studies show NASM as the cause of mastitis, some studies have demonstrated that NAS can reduce infection by S. aureus and other pathogens (Brouillette et al., Reference Brouillette, Goetz, Droppa-Almeida, Chamberland, Jacques and Malouin2022). In another study by Toledo-Silva et al., (Reference Toledo-Silva, de Souza, Mertens, Piepers, Haesebrouck and De Vliegher2021a, Reference Toledo-Silva, Nogueira De Souza, Piepers, Mertens, Haesebrouck and De Vliegher2021b), it was observed that NASM suppressed the biofilm dispersion of a strain of S. aureus. Some studies also point to a possible protective role of NASM, as described by Piepers et al. (Reference Piepers, Schukken, Passchyn and De Vliegher2013), who in comparing infected heifers with uninfected heifers, observed that heifers infected with NASM had fewer cases of clinical mastitis throughout lactation.
Some gram-positive bacteria can produce bacteriocins, inhibiting the growth of other bacterial strains. Bacteriocins produced by NASM include epidermin, hyicin 3682, nukacin ISK-1, nukacin 3299 and nukacin KQU-131. Staphylococcus epidermidis produces Tu 3298, Staphylococcus hyicus 3682, Staphylococcus warneri ISK-1, Staphylococcus simulans 3299, and Staphylococcus hominis KQU-131 (Yong et al., Reference Yong, Dykes and Choo2019), so it can be seen that there is potential for NASM to negatively affect the growth of pathogenic bacteria. In this context, Ferronatto et al. (Reference Ferronatto, Souza, Della Libera, De Vliegher, De Visscher, Piepers, Blagitz and Heinemann2019) used the cross-streaking method to test growth inhibition of mastitis-causing pathogens. Using this method, it is possible to determine if there was partial inhibition, total inhibition (no colony was observed), or no growth inhibition (same size and number of colonies as on the positive control plate). It was reported that all of the 19 NAS studied could inhibit growth partially, and in the case of S. chromogenes there was total inhibition. Another study showed the capacity of 38 strains of NAS to inhibit Listeria innocua growth, with the strain that most displayed this ability being S. quorum (Braem et al., Reference Braem, Stijlemans, Van Haken, De Vliegher, De Vuyst and Leroy2014).
De Vliegher et al. (Reference De Vliegher, Opsomer, Vanrolleghem, Devriese, Sampimon, Sol, Barkema, Haesebrouck and De Kruif2004) also studied how S. chromogenes could inhibit the growth of some bacteria, such as S. aureus, S. dysgalactiae and S. uberis, and the result was positive. The NAS studied inhibited the growth of these strains but not that of E. coli, which is typically expected, as the inhibitory effect of bacteriocins is usually more intense against phylogenetically related bacterial species (Ferronatto et al., Reference Ferronatto, Souza, Della Libera, De Vliegher, De Visscher, Piepers, Blagitz and Heinemann2019). Furthermore, the same authors in a previous study suggested that colonization of the teat apex by S. chromogenes in pre-calving heifers protected the udder against increased SCC in the early postpartum period (De Vliegher et al., Reference De Vliegher, Laevens, Devriese, Opsomer, Leroy, Barkema and De Kruif2003).
According to Isaac et al. (Reference Isaac, Bohl, Breser, Orellano, Conesa, Ferrero and Porporatto2017), NASM is a group of bacteria classified as either minor mastitis pathogens or commensal microbiota. A study showed that S. epidermidis, S. simulans, S. hominis, S. saprophyticus, and S. arlettae isolated from milk were able to produce antimicrobial compounds to inhibit the growth of some mastitis pathogens, including S. aureus (Nascimento et al., Reference Nascimento, Fagundes, Brito, Netto Dos Santos and Bastos2005). Beuckelaere et al. (Reference Beuckelaere, De Visscher, Souza, Meyer, Haesebrouck, Piepers and De Vliegher2021) observed a probiotic effect of S. chromogenes when colonizing dry quarters with this microorganism. As a result, the authors observed a shift to the Th1 response in late pregnancy and early lactation, increasing IgG2 concentration. In a study with a murine mastitis model, the authors observed that S. aureus intramammary colonization is reduced by exoproducts from S. chromogenes and S. simulans (Brouillette et al., Reference Brouillette, Goetz, Droppa-Almeida, Chamberland, Jacques and Malouin2022). Some NAS species (S. chromogenes, S. simulans and S. epidermidis) inhibited the growth of S. aureus and downregulated the expression of rnaIII, important molecule of the QS system (Toledo-Silva et al., Reference Toledo-Silva, de Souza, Mertens, Piepers, Haesebrouck and De Vliegher2021a, Reference Toledo-Silva, Nogueira De Souza, Piepers, Mertens, Haesebrouck and De Vliegher2021b). In addition, other studies point out that NASM protects the mammary quarters against IMI caused by primary pathogens (De Buck et al., Reference De Buck, Ha, Naushad, Nobrega, Luby, Middleton, De Vliegher and Barkema2021). oth characteristics of NASM, to cause or prevent mastitis, could be explained by differences among the species and intraspecies once the host determines the infection, the virulence of microorganisms and biofilm formation capacity (Brouillette et al., Reference Brouillette, Goetz, Droppa-Almeida, Chamberland, Jacques and Malouin2022). However, much still needs to be understood about these ratehr complex interactions within the microbiota.
One health and public health
The One Health approach conceptualizes that we cannot truly understand human, animal and environmental health by only addressing each in isolation. Because of this, we must understand the connections of these areas using holistic interpretations to face public health challenges. In other words, One Health focuses equally or more on the relationships between the factors in the system rather than on the individual-level factors themselves (Dalton et al., Reference Dalton, Rock, Carroll and Davis2020).
We can observe in Tables 1 and 2 NASM being isolated from food, clinical infections and environment, as well as their resistance to antibiotics. Several NASM strains from these studies (Kozioł-Montewka et al., Reference Kozioł-Montewka, Szczepanik, Baranowicz, Jóźwiak, Ksiazek and Kaczor2006; Hammad et al., Reference Hammad, Watanabe, Fujii and Shimamoto2012; Xu et al., Reference Xu, Liu, Chen, Liu, Tan, Shen and Zhang2019; Schnitt et al., Reference Schnitt, Lienen, Wichmann-Schauer and Tenhagen2021) hve been confirmed as MRNAS. Also, the topics about virulence and part of Table 1 showed the capacity of these species to produce toxins and form biofilm. All these components place NASM as a concern for public health.
Advances in molecular techniques allow us to relate strains from community and clinical infections, which helps us understand transmission among humans, animals and environments. The species most studied in Staphylococcus spp. is S. aureus and many studies have observed clones common in clinical infections, in animals or in the environment (Silva et al., Reference Silva, Guimarães, Manzi, Budri, Gómez-Sanz, Benito, Langoni, Rall and Torres2013, Reference Silva, Guimarães, Manzi, Júnior, Gómez-Sanz, Gómez, Langoni, Rall and Torres2014; Aires-de-Sousa, Reference Aires-de-Sousa2016; Zhou et al., Reference Zhou, Li, Shi, Wang and Yan2018). But, we also observe the same route for NAS in studies that demonstrate the presence of identical clones in hospitals and communities (Martínez-Meléndez et al., Reference Martínez-Meléndez, Morfín-Otero, Villarreal-Treviño, Camacho-Ortíz, González-González, Llaca-Díaz, Rodríguez-Noriega and Garza-González2016; Xu et al., Reference Xu, Liu, Chen, Liu, Tan, Shen and Zhang2019; Pinheiro-Hubinger et al., Reference Pinheiro-Hubinger, Riboli, Abraão, Pereira Franchi and Ribeiro de Souza da Cunha2021). So, the prevalence of NASM in the dairy chain and the increasing prevalence of resistance genes raises further concern. The prevalence of antimicrobial-resistant pathogens has increased while the approval of new drugs has decreased, which is a public health challenge. However, the number of antimicrobials used in veterinary medicine is immense and should be discussed since they select resistant microorganisms. Animals are the reservoirs of these organisms and can transmit them through food (Gohl et al., Reference Gohl, Vangay, Garbe, MacLean, Hauge, Becker, Gould, Clayton, Johnson, Hunter, Knights and Beckman2016; FDA, 2018).
According to the FDA, each year in the United States at least 2.8 million antibiotic-resistant infections occur, and more than 35 000 people die as a result. Therefore, agencies around the world are trying to minimize the harm caused by the incorrect use of antibiotics, for example, using them for purposes for which they are not intended (to treat diseases caused by viruses, fungi or yeasts), in addition to the frequent use of broad-spectrum antibiotics (Hildreth et al., Reference Hildreth, Burke and Glass2009). The FDA and the European Food Safety Authority (EFSA) are two organizations that have been playing a crucial role in the dissemination of vital information to preserve the effectiveness of currently available antimicrobial drugs and promote the development of new medical products that can help reduce the emergence and spread of antimicrobial resistant bacteria (EFSA, Reference Efsand; FDA, 2018).
Stevens et al. (Reference Stevens, Piepers, Supré and De Vliegher2018) analyzed the consumption of antimicrobials in dairy herds and its association with the diameters of the antimicrobial inhibition zone of NAS isolated from subclinical mastitis. They reported a high incidence of antimicrobial treatment for fourth generation cephalosporins, a class of antibiotics important for human health. Strains of MRNAS were identified in milk samples with mastitis in a study in Korea, and one of the strains persisted on one of the analyzed farms for more than two years (Kim et al., Reference Kim, Moon, Park, Kang, Na and Lim2019). The role of NAS as a cause of disease in humans has been described. Diseases such as endocarditis, tract infections, catheter-related sepsis, endophthalmitis, surgical site infections, peritonitis in patients with continuous ambulatory peritoneal dialysis and foreign body infections were due to infections caused by NAS (Piette and Verschraegen, Reference Piette and Verschraegen2009). Approximately 55–75% of nosocomial isolates are resistant to methicillin, and NASM is responsible for about 30% of all nosocomial bloodstream infections (Piette and Verschraegen, Reference Piette and Verschraegen2009).
In the context of One Health and mastitis, there is a solid recommendation for reducing antimicrobial use. Taking into acount that milk with antibiotic residues does not enter the supply chain, as it is withdrawn at source, this recommendation emphasizes the need to periodically search for antimicrobial residues in raw milk on farm. Combined with microbiome analysis and research on antimicrobial residues, we have the somatic cell count, which acts as an indicator of the health of the mammary gland, helping to reveal the situation of the dairy herd within the context of health (Gohl et al., Reference Gohl, Vangay, Garbe, MacLean, Hauge, Becker, Gould, Clayton, Johnson, Hunter, Knights and Beckman2016). When milk contains residues, this indicates that it comes from unhealthy animals, so it can cause foodborne pathogens and be a potential allergen for consumers. This concern is even more significant in developing countries, where there is no supervision on using antimicrobials in animals or the opportunity to test milk and meat for drug residues. The presence of adulterating antimicrobials in milk can have a profound impact on the health of babies and children in regions that already face high rates of diarrheal diseases and malnutrition. Several studies have shown a relationship between the intestinal microbiome, health status and disease. Therefore, ensuring that milk does not contain antimicrobial residues significantly impacts public health and producers' economic livelihood (Mohan et al., Reference Mohan, Koebnick, Schildt, Mueller, Radke and Blaut2008; Gordon et al., Reference Gordon, Dewey, Mills and Medzhitov2012; Garcia et al., Reference Garcia, Osburn and Cullor2019). Since NASM is a large group that contains species with different levels of pathogenicity, virulence, resistance profile and host adaptation, analyses of the genetic profile of these species are necessary to understand the role of each NAS species in bovine mastitis (De Buck et al., Reference De Buck, Ha, Naushad, Nobrega, Luby, Middleton, De Vliegher and Barkema2021). Thus, the One Health approach is also required when we are specifically talking about NAS.
This article gathers evidence that NASM has the potential to cause mastitis, and that all of this can cause public health problems since there is a lack of specific studies and strategies to combat these pathogens. Even with this evidence, NASM is still overlooked as causing disease in both humans and animals. With all these points raised, we must apply a One Health approach to address this critical public health problem, including implementing good antimicrobial administration practices in human and veterinary health settings to delay the development of resistance and prolong the life of useful antimicrobials (FDA, 2018).
In conclusion, since it is an important food at all stages of life, milk consumption increases each year along with population growth. We know that NASM mastitis can harm the health and welfare of animals and, consequently, human health, either by food poisoning or by the recurrent use of antimicrobials to treat the disease, which can leave residues in the milk that will be consumed in different ways. Studies related to NASM mastitis are extremely important to learn more about the role of these microorganisms in the disease and how their virulence factors can make treatment difficult. Questions that need more concrete answers are why some NASM strains that cause mastitis appear to have a protective role against other microorganisms and why some strains can harm milk production and animals.
In addition, the increased resistance of these microorganisms to antimicrobials is worrying, requiring the implementation of residue monitoring in milk and studies of NASM antimicrobial resistance used to combat mastitis. Thus, a holistic vision that encompasses safer food and solutions to improve animal, human and ecosystem health seems to be the best alternative.
Acknowledgements
This work was carried out with the support of the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior – Brasil (CAPES) – Finance Code 001 and São Paulo Research Foundation – FAPESP, grant #2019/17308-4.




