Cluster of differentiation 4 (CD4) is a single-pass transmembrane glycoprotein playing an important function in the interaction between T lymphocytes and antigen- carrying cells (König and Zhou, Reference König and Zhou2004). In Bos taurus, CD4 gene is located on chromosome 5 and two isoforms X1 (GenBank Accession Number: XM_024991416) and X2 (XM_024991417) are transcripted. The coded proteins are made of 455 (XP_024847184) and 395 (XP_024847186) amino acids respectively, due to splicing of exon 7 and variation in the reading frame.
The influence of some variants in the CD4 genomic sequence is already known in humans for their ability to increase susceptibility to HIV infection (Oyugi et al., Reference Oyugi, Vouriot, Alimonti, Wayne, Luo, Land, Ao, Yao, Sekaly, Elliott, Simonsen, Ball, Jaoko, Kimani, Plummer and Fowke2009) or for their association to the susceptibility and severity of rheumatoid arthritis (Hussein et al., Reference Hussein, El Tarhouny, Mohamed, El-Shal, Abul-Saoud and Abdo2012). Also, in the animal field recent researches have highlighted an important role of CD4 in immunity. In particular, Wang et al. (Reference Wang, Zhang, He, Ma, Fan, Wang, Zhang, Sun, Zhang, Wang, Song and Yu2013) found the CD4 gene to be linked to mastitis resistance in dairy cows. He et al. (Reference He, Chu, Ma, Wang, Zhang, Sun, Zhang, Yu and Zhang2011) highlighted a significant association between SNPs within CD4 gene with mastitis indicators and milk production traits. Usman et al. (Reference Usman, Yu, Zhai, Liu, Wang and Wang2016) showed a significant association between some SNPs of the CD4 gene (between exon 2 and intron 6) and the percentage of fat in milk. Recently, Zeb et al. (Reference Zeb, Ali, Niaz, Rasheed, Khattak, Khan, Wang and Usman2020) showed that four SNPs between exon 2 and exon 4 of the CD4 gene were significantly associated with clinical mastitis incidence and with annual milk yield. Grandoni et al. (Reference Grandoni, Napolitano, Scatà and De Matteis2020) identified in two Italian breeds (Friesian and Simmental) and their crosses two new missense mutations in the exon 5: g.103637203 G > T and g.103637191 A > C. The mutations were involved in lack or reduced labeling of anti-CD4 monoclonal antibody clone CC8 detected by flow cytometry assay.
In the present study, the entire CD4 gene sequences of Simmental sires were investigated in order to mevaluate the effects of genomic variants on their deregressed multiple across country evaluation (MACE) of estimated breeding values (EBV) indexes for milk, fat and protein yields, and for somatic cell score (SCS). This index allows one to obtain the genetic evaluations of the reproducers, elaborated by the various countries, in a single international evaluation. Using the MACE model, Interbull (International Bull Evaluation Service) evaluates each trait considering the estimates of the genetic correlations between sires in the different countries. This paper documents the presence of a substantial number of polymorphisms in the CD4 gene and their significant associations on milk and protein indexes.
Material and methods
Data used in this study came from the Italian Simmental Breeders Association (ANAPRI). The Association provided for the complete genomic sequences and the deregressed MACE EBV indexes of 34 Simmental sires of different origins (21 Austrian, 7 Swiss, 4 German and 2 Italian) and born between 1969 and 2006. The sires had an average of 6,316 daughters with a minimum of 95 daughters and a maximum of 42 657. Indexes concerning yields of milk, fat and protein as well as SCS and their genetic basis referred to the years 2013–2015, with an update happening every five weeks.
Genetic variants in the CD4 gene
In order to investigate the presence of polymorphisms in the coding and regulatory regions of CD4 gene, we used the complementary sequence of the genomic region of chromosome 5 in Dominette's Bos taurus (GeneBank, Accession Number: NC_037332). This sequence ranges between the nucleotides g.103631000…103660000 (29.000 bp) and it contains both the coding and regulatory regions of the bovine CD4 gene.
Genome-wide sequencing of each of the 34 bulls was uploaded to Galaxy server at https://usegalaxy.eu (Version 2.3.4.3). Bowtie 2, tool of the Galaxy software, was used to map reads of the 34 bulls on the region of chromosome 5 including the CD4 gene, after assessing the quality of the raw reads through Galaxy's FastQC tool.
Bioinformatic analysis
Bioinformatic analysis was performed using the MATCHTM programme (Kel et al., Reference Kel, Gossling, Reuter, Cheremushkin, Kel-Margoulis and Wingender2003) in the TRANSFAC® Professional 10.2 http://www.biobaseinternational.com/ (Matys et al., Reference Matys, Kel-Margoulis, Fricke, Liebich, Land, Barre-Dirrie, Reuter, Chekmenev, Krull, Hornischer, Voss, Stegmaier, Lewicki-Potapov, Saxel, Kel and Wingender2006). A profile search was performed for transcription factor binding sites in the sequenced DNA fragments, focusing on the areas where SNPs had been detected.
The ‘vertebrate binding’ and the ‘only high quality’ matrix options were used in the profile selection, with cut-offs for core and matrix similarity set to 0.75 and 0.7 respectively, without any changes in other options.
Statistical analysis
On each SNP site, the χ2 test for deviation from Hardy-Weinberg equilibrium, along with expected and observed heterozygosity, polymorphism information content (PIC) and linkage disequilibrium (LD) were calculated using the algorithms provided for by the SAS software 9.4 (ALLELE procedure).
The associations between SNPs and traits were analysed using the general linear model (GLM procedure of SAS software 9.4): Y ik = μ + G i + e ik where Y ik is the phenotype, μ was the overall population mean for the trait, G i is the fixed effect of the genotype at the SNP (i = AA, AB, BB) and e ik is the random error. Significance of the SNP and least-squares means were determined using the Student's t-test in GLM procedure.
The association between haplotypes and traits were analysed by a GLM on each haplotype by inserting the number of copies as a fixed: Y ik = μ + H i + e ik, where Y ik is the phenotype, μ is the overall population mean for the trait, H i is the fixed effect of the haplotype [i = 0, individuals with no haplotype; 1, individuals with one copy only (heterozygotes); 2, individuals homozygous for that haplotype] and e ik is the random error. Significance of the haplotype and least-squares means were determined using the Student's t-test in GLM procedure.
Results
The comparison of two already known sequences of bovine mRNA (transcript variant X1, NCBI accession: XM_024991416; transcript variant X2, NCBI accession: XM_024991417) with the DNA fragment of 29 000 bp deriving from the genomic sequence of the Dominette bovine, suggested the use of this DNA fragment as reference basis for the study of possible variants in the CD4 gene. In fact, this genomic fragment contains both the coding and regulatory parts of the two transcript isoforms. The X1 and X2 variants differ in two respects: a different 5′UTR and the lack of exon 7 in variant X2. While the 5′UTR of the X1 variant ranges between the nucleotides g.103654819 …103654611, the same UTR in the X2 variant ranges between g.103651793 …103651649 of the bovine sequence (NC_037332). Furthermore, the splicing of exon 7 in the X2 variant involves a modification in the reading of the nucleotide sequence with the formation of a stop codon (TAG) in position g.103633422 and the consequent reduction of the number of amino acids in the protein isoform (395aa – in X2 – XP_024847186 as compared to the 455aa in X1 – XP_024847184).
Genome-wide sequencing analysis of 34 bulls uploaded to the Galaxy server showed that the number of raw reads per bull on average is approximately 181 million, for both forward and reverse sequencing. The quality control of these reads, produced by the FastQC report, highlights a good quality of the sequences expressed by an average Phred Score value of 38, remembering that the minimum value for a good sequence is 20 (Kwong et al., Reference Kwong, McCallum, Sintchenko and Howden2015). Aligning the reads with the Bowtie2 tool, on the 29 000 bp genomic sequence of the CD4 gene, showed an average of 6.2 million reads for both forward and reverse bull sequencing. Furthermore, only genomic variants with a Phred Quality Score > 40 were considered.
Diversity analysis of CD4 SNPs
Thirty-one SNPs were identified in the described genomic sequence (Table 1) sixteen of them causing sequence variations in coding or regulatory regions of the CD4 gene (SNPs: 1, 2, 11 ÷ 17, 21 ÷ 24, 27 and 30 ÷ 31), and the others located in regions not expected to affect gene functionality. Among those positioned in coding regions, SNPs 11 ÷ 17 (exon 2), 22 (exon 4), 23 and 24 (exon 5), produce changes in the amino acid sequence, while the other two are synonymous (21 of exon 3 and 27 of exon 6). Three SNPs were newly found in the CD4 gene, namely: SNP 19 and 20, identified in intron 2 and SNP 30 present in the 3′-UTR regulatory region.
Table 1. Information on genetic variants of the CD4 gene in Simmental breed.
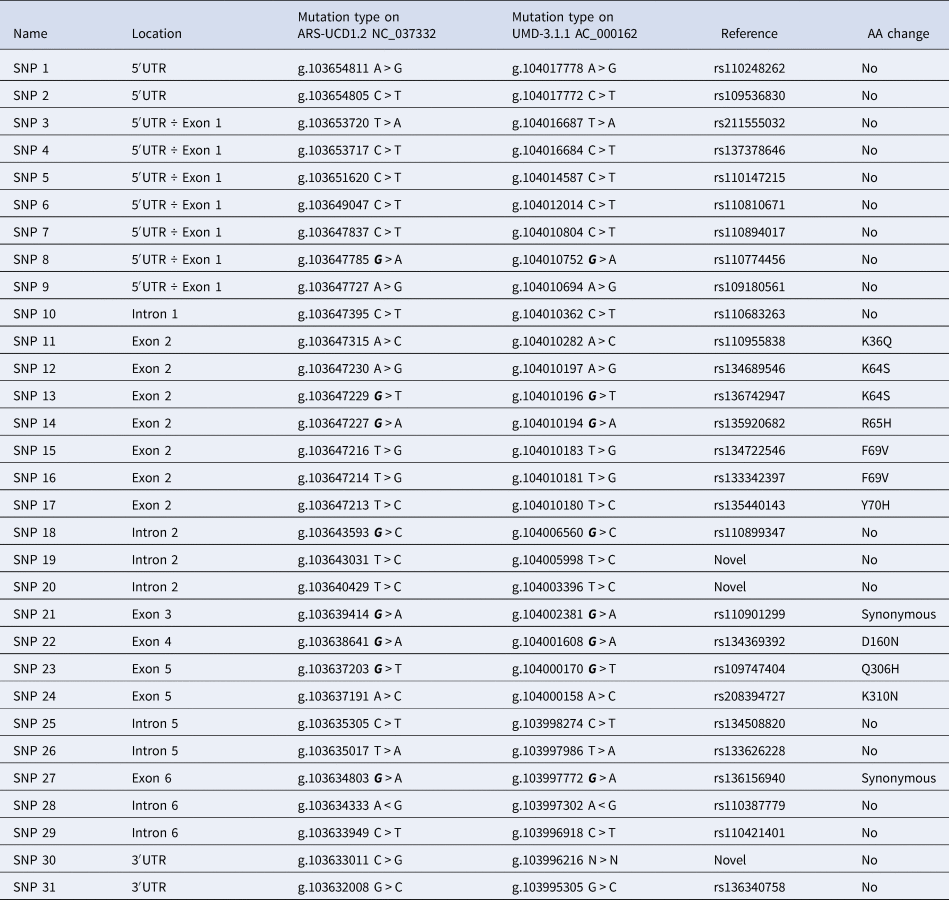
Most of the SNPs are informative markers (online Supplementary Table S1), with a PIC value over 0.20. Only genotypes at SNPs 7 and 19 were not in Hardy-Weinberg equilibrium (χ2 = 21.19 and 14.74, respectively; P < 0.0001), possibly due to some sort of selection.
Correlations among SNPs are reported in online Supplementary Table S2. Sixteen SNPs (1, 9 ÷ 18, 21, 22, 25, 28 and 29), one half of the total number, were 100% in linkage disequilibrium (LD = 1). Out of these, one was chosen for the association analysis based on its location in the coding region of the gene and because it had already been used by other authors (Usman et al., Reference Usman, Yu, Zhai, Liu, Wang and Wang2016; Zeb et al., Reference Zeb, Ali, Niaz, Rasheed, Khattak, Khan, Wang and Usman2020) in association tests in dairy cattle: SNP 11 (rs110955838) in exon 2. Ten more SNPs with LD < 0.9: SNP 2 (rs109536830), SNP 3 (rs211555032), SNP 5 (rs110147215), SNP 7 (rs110894017), SNP 8 (rs110774456), SNP 19 (novel), SNP 20 (novel), SNP 27 (rs136156940), SNP 30 (novel), and SNP 31 (rs136340758) were processed for the estimation of their associations with phenotypes (deregressed MACE EBV indexes of milk, fat and protein yields, and SCS) as function of their genotype.
Association analysis of each SNP on milk indexes
Table 2 reports the association of the genotypes at the 11 SNP loci with the indexes on milk, fat and proteins yields, and SCS. The association analysis indicated that only four of the eleven chosen markers had an effect on some of the considered traits. In particular, SNPs: 3, 8, 11, and 19, exhibited a significant effect (P < 0.05) on protein yield index; SNP 19 exhibited a significant effect (P < 0.05) also on milk yield, while SNP 11 showed a numerical but non-significant difference (P > 0.05) on the same trait.
Table 2. Association of SNPs of the CD4 gene with the deregressed multiple across country evaluation of estimated breeding values indexes of some milk-related traits in Simmental breed.*
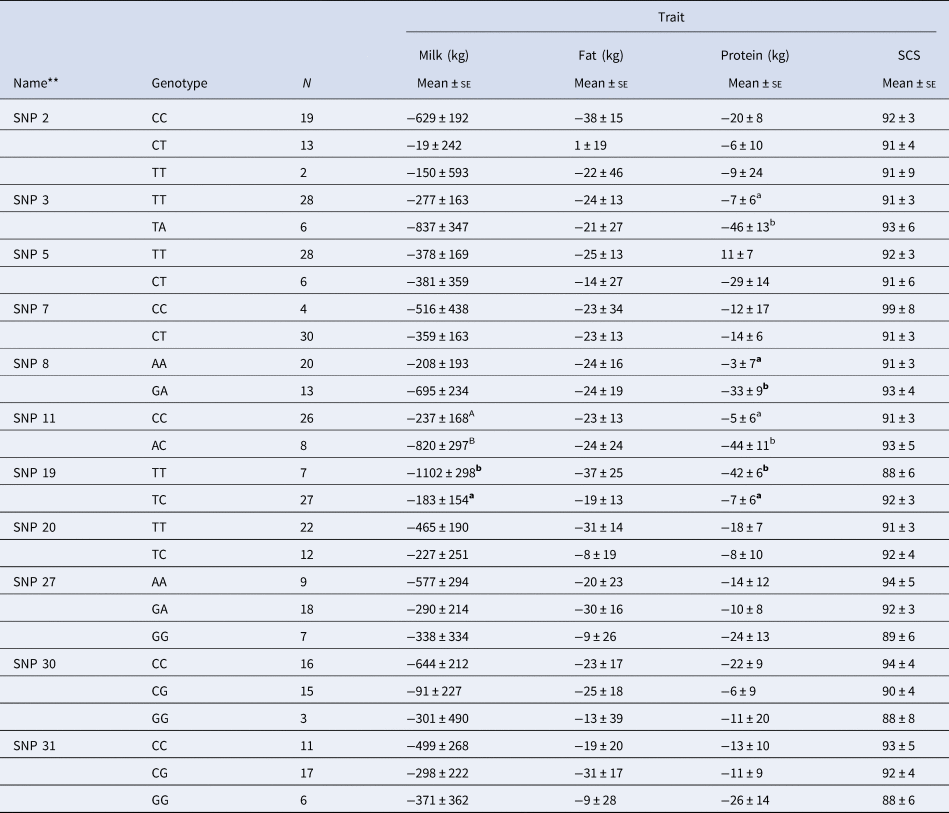
* Within each group, values with differing superscript letters mean a significant difference (a, b = P < 0.05) or numerical but non-significant difference (A, B = P < 0.10).
** Novel SNPs are marked in bold.
Bioinformatic analysis on the detected SNPs
The identification of SNPs affecting the considered indexes even if located in non-coding or regulatory regions (SNPs: 3, 8 and 19) led us to evaluate the existence of transcription factor sites near them.
Transcription factors identified in TRANFAC database were present in the regions of the CD4 gene modified by our markers. SNP 3 T > A was located into the ‘core’ of the consensus sequence (ggT/AGACAcctgt, g.103653722…711 on NC_037332) of binding site for transcription factors CREB (cAMP Response Element Binding Protein). The novel SNP 19 T > C was located into the ‘core’ of the consensus sequence (tttGATCT/Cct, g.103643038…029 on NC_037332) for GATA-3 (GATA-binding protein 3). No transcription factors were detected in the sequence affected by the marker SNP 8.
Association analysis of haplotypes on milk indexes
As reported in Table 2, 4 out of 11 analysed SNPs (3, 8, 11 and 19) produced significant effects on milk related indexes. To evaluate the existence of haplotypic effects, combinations among the four genomic variants were investigated.
Haplotype make-up and frequencies are listed in Table 3. Six different haplotypic alleles were identified, but only four of them were frequent enough to allow for an evaluation of any haplotypic effect (at least six copies in the sample examined): Hap1, Hap2, Hap3, and Hap6.
Table 3. Information on haplotypic alleles of the CD4 gene in Simmental breed.
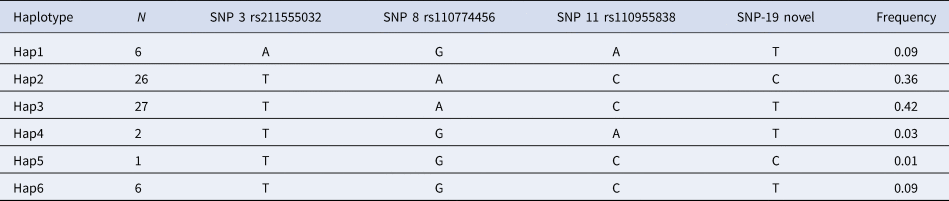

The analysis of the associations between the selected haplotypes in the CD4 gene with milk related indexes (Table 4) showed that bulls with haplotype Hap2 (T-A-C-C) had better indexes for milk and protein yields (P < 0.05), whereas the presence of the Hap1 haplotype (A-G-A-T) caused a significant decrease of the index for protein yield (P < 0.05). Frequencies of the two alleles Hap1 and Hap2 (9 and 36% respectively) make them of particular interest.
Table 4. Association of the haplotypic alleles of the CD4 gene with the deregressed multiple across country evaluation of estimated breeding values indexes of some milk traits in Simmental breed*
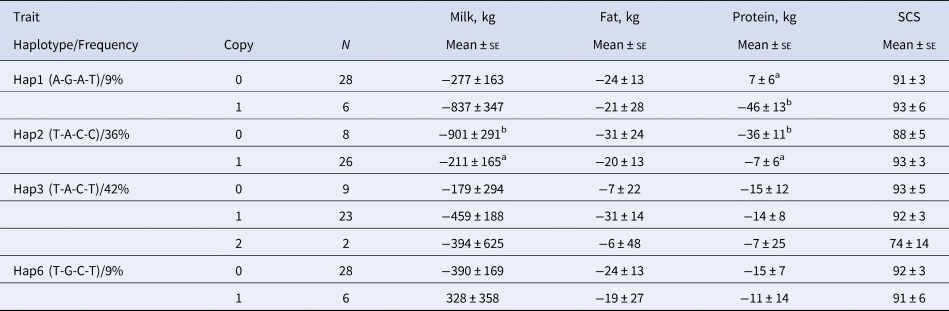
* Within each group, values with differing superscript differ significantly, P < 0.05 or greater.
Discussion
This study documents the occurrence of several polymorphisms in the genomic sequence of the entire CD4 gene in a sample of Simmental genomic sires. Many of these polymorphisms, as already reported by other authors (Usman et al., Reference Usman, Yu, Zhai, Liu, Wang and Wang2016; Zeb et al., Reference Zeb, Ali, Niaz, Rasheed, Khattak, Khan, Wang and Usman2020), were in perfect linkage disequilibrium in the surveyed 34 genomes. In the present work we investigated the possible effect of genomic variants in CD4 on the deregressed MACE EBV indexes for SCS, an indicator of udder health status and for milk quality and quantity.
The identified variants insist on coding regions (variants of exons 2, 3, 4, 5 and 6), in regulatory regions (5′ and 3′ UTR) and also within intronic sequences. Attention to these last two areas of the CD4 gene revealed the presence of sites for transcription factors.
SNP 3 (rs211555032) located into the ‘core’ of the consensus sequence of binding site for CREB, determined a significant and negative effect on protein yield index. CREB is a transcription factor that regulates diverse cellular responses, including proliferation, survival, and differentiation. Its activation is required for the generation and maintenance of regulatory T cells (Wen et al., Reference Wen, Sakamoto and Miller2010). Furthermore, as reported by Pegolo et al. (Reference Pegolo, Mach, Ramayo-Caldas, Schiavon, Bittante and Cecchinato2018) the transcription factor CREB is known to regulate milk protein synthesis.
In our sample of Simmental sires the SNP 8 (rs110774456) highlights a significant and negative effect on the protein yield index. No effect was detected on the other traits, in particular for the somatic cell score.
The analysis of the genetic variants of the CD4 gene highlights that SNP 11 (rs110955838) is in very close linkage disequilibrium with most of the variants present in the coding and regulatory areas of the gene. This is also evident from the results obtained by other authors in the analysis of the association between some SNPs of CD4 and production traits in dairy cattle (Usman et al., Reference Usman, Yu, Zhai, Liu, Wang and Wang2016; Zeb et al., Reference Zeb, Ali, Niaz, Rasheed, Khattak, Khan, Wang and Usman2020). He et al. (Reference He, Chu, Ma, Wang, Zhang, Sun, Zhang, Yu and Zhang2011) reported a significant association between the SNP g.13598 C > T of the CD4 gene in intron 6, which corresponds to the SNP 29 (rs110421401) and which is in perfect linkage disequilibrium with the SNP 11, with the production traits of milk and protein yield in Chinese Holsteins, in agreement with our results on Simmental sires. In their study, the authors also highlighted a significant effect on the somatic cell score trait, which is not highlighted in our data. Other studies reported association of SNPs in CD4 gene with fat percentage but not with SCS (Usman et al., Reference Usman, Yu, Zhai, Liu, Wang and Wang2016), as well as Zeb et al. (Reference Zeb, Ali, Niaz, Rasheed, Khattak, Khan, Wang and Usman2020) that describe a significant association with the frequency of clinical mastitis but not with SCS. The same authors identify possible reasons for these discrepancies in the differences in the breeds of cattle, the environment conditions, and population structure.
The novel SNP 19 was located in the ‘core’ of the consensus sequence of binding site for the transcription factor GATA-3 and determined a significant and positive effect on milk and protein yield indexes. The zinc finger transcription factors GATA-3 was defined as a master regulator of the mammary gland (Tong and Hotamisligil, Reference Tong and Hotamisligil2007): its expression is required for normal embryonic mammary development, while in the postnatal mammary gland, GATA-3 is essential for the formation of the ductal tree and for differentiation of the epithelial cells (Kouros-Mehr et al., Reference Kouros-Mehr, Slorach, Sternlicht and Werb2006; Tong and Hotamisligil, Reference Tong and Hotamisligil2007; Kinoshita et al., Reference Kinoshita, Yoshizawa, Emoto, Yuki, Yuri, Shikata and Tsubura2014). Gata-3 expression is also essential for early T cell development in the thymus and for differentiation of naïve CD4+ T cells into committed T helper type 2 cells (Skapenko et al., Reference Skapenko, Leipe, Niesner, Devriendt, Beetz, Radbruch, Kalden, Lipsky and Schulze-Koops2004; Tindemans et al., Reference Tindemans, Serafini, Di Santo and Hendriks2014).
The haplotypic approach allows us to gain a systemic view of the probable role of the CD4 gene in the determinism of livestock traits. This study, within the limits of our sample, highlighted the association of some of the newly found haplotypes with milk-related deregressed MACE EBV indexes in Simmental breed. Therefore, they could be used as genetic markers in future breeding programs. For example, it should be possible to increase the frequency of haplotype Hap2 (T-A-C-C) in the population, if the selection goals included milk quality and yield improvement. Protein yield appear to be significantly depressed by Hap 1 (A-G-A-T). As it can be easily seen, the two haplotypes differ in all four SNPs contributing to their identification. This is important as an indirect confirmation of the results from SNP association analysis.
Although the role of CD4 gene during inflammation is well documented (Oyugi et al., Reference Oyugi, Vouriot, Alimonti, Wayne, Luo, Land, Ao, Yao, Sekaly, Elliott, Simonsen, Ball, Jaoko, Kimani, Plummer and Fowke2009; Usman et al., Reference Usman, Yu, Zhai, Liu, Wang and Wang2016), little is known about its possible association with production traits. This work implements knowledge on CD4 genomic variability through a ‘candidate gene’ approach, highlighting three novel variants. These new SNPs, together with the many other ones previously identified by other authors, allowed us to identify new haplotypes considering the entire gene sequence. The significant association of CD4 haplotypes with milk and protein indexes observed in this study supports the hypothesis of considering this gene as a candidate for the improvement of milk-related traits in Simmental breed.
In conclusion, we investigated the sequence of the entire CD4 gene in Simmental sires to evaluate the effects of genomic variants on the indexes of the bulls for milk, fat and protein yields, as well as somatic cell score. Only four of thirty-one identified SNPs influenced these traits. To evaluate the existence of haplotypic effects, combinations among these four were investigated and six different haplotypic alleles were identified, of which four were frequent enough to allow for a full evaluation: Hap1, Hap2, Hap3 and Hap6. The analysis of associations between the selected haplotypes in the CD4 gene with milk related indexes showed that bulls with Hap2 (T-A-C-C) had better indexes for milk and protein yields whereas the presence of the Hap1 haplotype (A-G-A-T) caused a significant decrease of the index for protein yield. Frequencies of the two alleles Hap1 and Hap2 (9 and 36% respectively) make them of interest for their possible inclusion in breeding schemes and support the hypothesis of considering this gene as a candidate for the improvement of milk-related traits in the Simmental breed.
Supplementary material
The supplementary material for this article can be found at https://doi.org/10.1017/S0022029921000832






